Tracer v1.6 model comparison very SLOW
1,095 views
Skip to first unread message
Qiyun Zhu
Jan 10, 2014, 7:02:56 PM1/10/14
to beast...@googlegroups.com
Hi all,
I tried the Tracer version 1.6 to perform model comparison. Unlike Tracer 1.5's "Bayes Factor", which runs decently fast, the new Tracer's "Model comparison" (HME and AICM) runs incredibly slow, and technically never ends. My dataset has a considerable size (250 taxa, 10000 characters, 14 partitions, 25 million generations). I feel (just feel) that as the number of bootstrap relication increases, the computation time increase at an exponential pace (not linear, as I commonly understand). I hereby report this issue to the community and wish someone could tell me "hey, the way you run Tracer is not right!". Thanks!
Best,
Qiyun
I tried the Tracer version 1.6 to perform model comparison. Unlike Tracer 1.5's "Bayes Factor", which runs decently fast, the new Tracer's "Model comparison" (HME and AICM) runs incredibly slow, and technically never ends. My dataset has a considerable size (250 taxa, 10000 characters, 14 partitions, 25 million generations). I feel (just feel) that as the number of bootstrap relication increases, the computation time increase at an exponential pace (not linear, as I commonly understand). I hereby report this issue to the community and wish someone could tell me "hey, the way you run Tracer is not right!". Thanks!
Best,
Qiyun
Guy Baele
Jan 11, 2014, 2:31:24 PM1/11/14
to beast...@googlegroups.com
Hi Qiyun,
Op zaterdag 11 januari 2014 01:02:56 UTC+1 schreef Qiyun Zhu:
I'm not sure why Tracer 1.6 would be slower in running those bootstraps. However, I'd like to point out that performing model comparison using HME/sHME and AICM is simply not a good idea as we've shown in a number of papers where all the analyses were done in BEAST (Baele et al. MBE 2012, MBE 2013, Bioinformatics 2013). And running those bootstrap replicates is also something I'd like to discourage for inaccurate estimators. You should try switching to path sampling (PS) and stepping-stone sampling (SS) to do your model comparisons.
Best regards,
Guy
Op zaterdag 11 januari 2014 01:02:56 UTC+1 schreef Qiyun Zhu:
Qiyun Zhu
Jan 12, 2014, 12:08:42 PM1/12/14
to beast...@googlegroups.com
Hello Guy,
Thanks for your kind reply! I knew your studies and I have tried PS and SS. Since they are implemented in BEAUti 1.8.0, it's pretty straightforward to try. However my concern is the computation burden. The default chain length (10M) and number of steps (100) makes the task consuming 100 times as much time as the original analysis, and is therefore technically impossible on our poor little server. Therefore I had to reduce chain length to 100K in order to control the running time within an acceptable range. But I fear this is not accurate enough. Could you please recommend me a parameter setting that balances computation expense and accuracy? Thank you!
Best,
Qiyun
Qiyun Zhu
Jan 12, 2014, 12:58:39 PM1/12/14
to beast...@googlegroups.com
Hello Guy,
Besides, your paper mentioned burn-in in PS and SS analysis: "PS and SS were run for an initial burn-in of 2.5 million iterations, after which 64 power posteriors were run for 275 thousand iterations each, discarding the first 25 thousand iterations as the burn-in."
However, in BEAST interface / example / tutorial I did not find anywhere that I can specify a burn-in. So I assume I don't have to manually specify burn-in in PS and SS. Is that correct?
Besides, your paper mentioned burn-in in PS and SS analysis: "PS and SS were run for an initial burn-in of 2.5 million iterations, after which 64 power posteriors were run for 275 thousand iterations each, discarding the first 25 thousand iterations as the burn-in."
However, in BEAST interface / example / tutorial I did not find anywhere that I can specify a burn-in. So I assume I don't have to manually specify burn-in in PS and SS. Is that correct?
Best,
Qiyun
On Saturday, January 11, 2014 2:31:24 PM UTC-5, Guy Baele wrote:
Guy Baele
Jan 13, 2014, 8:11:09 AM1/13/14
to beast...@googlegroups.com
Hi Qiyun,
Op zondag 12 januari 2014 18:08:42 UTC+1 schreef Qiyun Zhu:
There must have been a small glitch in setting the default chain length, it should be 1M (I've now changed this in the source code). But yes, that would still make your task 10 times as challenging as the original analysis. There are no settings (except possibly extremely demanding ones) that will work for every data set. But the more iterations you need to run your original analysis, the more demanding the marginal likelihood estimation will be. I fear that a chain length of 100K will indeed not be accurate enough. Could you try running 50 path steps with 500K each?
Best regards,
Guy
Op zondag 12 januari 2014 18:08:42 UTC+1 schreef Qiyun Zhu:
Guy Baele
Jan 13, 2014, 8:12:54 AM1/13/14
to beast...@googlegroups.com
Hi Qiyun,
Op zondag 12 januari 2014 18:58:39 UTC+1 schreef Qiyun Zhu:
The burn-in for each power posterior, when performing marginal likelihood estimation using PS/SS, is set at 10% by default, which is what I used in that paper. The initial burn-in of 2.5 million iterations that is mentioned refers to the preceding chain that was run to explore the posterior, which is the chain length defined in the mcmc block.
Best regards,
Guy
Op zondag 12 januari 2014 18:58:39 UTC+1 schreef Qiyun Zhu:
Qiyun Zhu
Jan 13, 2014, 9:20:30 AM1/13/14
to beast...@googlegroups.com
Hello Guy,
Thank you so much for your explanations, which for sure extend our understandings of the latest functions of BEAST. Yes I can try 500K x 50 steps, which probably will take less than week for one run. I will post my results once done.
By the way, how many replicates should I do per topological hypothesis?
Based on the AICM and SS tests I already did, I found that the results still flucturate a lot. The same topological hypothesis can yield log marginal likelihood values that differ by dozens or even hundreds among replicates. For two conflicting hypotheses, the worse scenario indeed yields a lower average SS value than the better one, however the SS of some replicates from the worse group may be even higher than that of some replicates from the better group. I looked at the traces of likelihood in Tracer and noticed that replicates from the same group tend to stabilize at slightly different levels. I guess this may be because the MCMC got stuck at some sub-optimal pits in the tree space. Therefore I am thinking if I should place more strict topological constraints, for example, simply give BEAST a starting tree and fix the tree topology to it. How do you think of this idea?
Best,
Qiyun
Thank you so much for your explanations, which for sure extend our understandings of the latest functions of BEAST. Yes I can try 500K x 50 steps, which probably will take less than week for one run. I will post my results once done.
By the way, how many replicates should I do per topological hypothesis?
Based on the AICM and SS tests I already did, I found that the results still flucturate a lot. The same topological hypothesis can yield log marginal likelihood values that differ by dozens or even hundreds among replicates. For two conflicting hypotheses, the worse scenario indeed yields a lower average SS value than the better one, however the SS of some replicates from the worse group may be even higher than that of some replicates from the better group. I looked at the traces of likelihood in Tracer and noticed that replicates from the same group tend to stabilize at slightly different levels. I guess this may be because the MCMC got stuck at some sub-optimal pits in the tree space. Therefore I am thinking if I should place more strict topological constraints, for example, simply give BEAST a starting tree and fix the tree topology to it. How do you think of this idea?
Best,
Qiyun
Guy Baele
Jan 14, 2014, 7:14:02 AM1/14/14
to beast...@googlegroups.com
Hi Qiyun,
Op maandag 13 januari 2014 15:20:30 UTC+1 schreef Qiyun Zhu:
Running multiple replicates only makes sense when the marginal likelihood estimates have converged close enough to the true value.
If not, then you're merely obtaining a mean marginal likelihood (with some standard deviation) that could be far off the correct result.
If not, then you're merely obtaining a mean marginal likelihood (with some standard deviation) that could be far off the correct result.
When you are positive that your settings are demanding enough given your data set, then running several replicates may point out potential convergence problems.
Not only for a standard MCMC analysis, but also for marginal likelihood estimations, each chain needs to run long enough to ensure convergence.
This could be what you are seeing here. It could be due to the fact that tree space is vast and difficult to explore or it might be because of another parameter.
If you're interested in comparing a discrete set of topologies, then of course, that's something you can do.
Best regards,
Guy
Op maandag 13 januari 2014 15:20:30 UTC+1 schreef Qiyun Zhu:
Qiyun Zhu
Jan 14, 2014, 1:13:03 PM1/14/14
to beast...@googlegroups.com
Hello Guy,
I appreciate your kind explanation from the perspective of designer and developer. Parameters have already been carefully optimized so that none of them looks odd in Tracer. In my previous experience, the analysis goes smooth without constraints, but when I force a less optimal topological constraint, the traces begin to go wildly. Perheps increasing number of iterations does help to solve this problem. I will probably try to double the number and test again.
Best,
Qiyun
I appreciate your kind explanation from the perspective of designer and developer. Parameters have already been carefully optimized so that none of them looks odd in Tracer. In my previous experience, the analysis goes smooth without constraints, but when I force a less optimal topological constraint, the traces begin to go wildly. Perheps increasing number of iterations does help to solve this problem. I will probably try to double the number and test again.
Best,
Qiyun
Qiyun Zhu
Jan 19, 2014, 4:59:19 PM1/19/14
to beast...@googlegroups.com
Hello Guy,
I have tested a longer MCMC chain with 500K x50 steps as you recommended. I found that 500 x50 is not that computation-expensive, however, after ~40 steps, the program kept saying "underflow ... rescaling ...", and became much slower. The last 10 steps acutally took more than 1/3 of total computation time. I am not sure if it's normal. Anyway, doing more steps seem to be more difficult. I uploaded the Tracer screenshot here. Could you please comment whether this trace looks reasonable? Thank you!
Best,
Qiyun
I have tested a longer MCMC chain with 500K x50 steps as you recommended. I found that 500 x50 is not that computation-expensive, however, after ~40 steps, the program kept saying "underflow ... rescaling ...", and became much slower. The last 10 steps acutally took more than 1/3 of total computation time. I am not sure if it's normal. Anyway, doing more steps seem to be more difficult. I uploaded the Tracer screenshot here. Could you please comment whether this trace looks reasonable? Thank you!
Best,
Qiyun
Guy Baele
Jan 20, 2014, 4:43:32 AM1/20/14
to beast...@googlegroups.com
Hi Qiyun,
Op zondag 19 januari 2014 22:59:19 UTC+1 schreef Qiyun Zhu:
You're not really supposed to load the likelihood samples from the marginal likelihood computations into Tracer (even though you can).
What you're seeing is hence normal, the likelihoods on average become lower as you're getting closer to the prior.
About the computation slowing down the closer you get to the prior, that also happens from time to time, depending on the data set and the models you're using.
Are you using the dynamic scaling option in BEAGLE (should be the default option I think) ?
This should make things considerably faster, but be sure to double check your results if you get a lot of underflow warnings.
Best regards,
Guy
Op zondag 19 januari 2014 22:59:19 UTC+1 schreef Qiyun Zhu:
Qiyun Zhu
Jan 21, 2014, 4:42:32 PM1/21/14
to beast...@googlegroups.com
Hello Guy,
Thanks for the explanation. Yes I used dynamic scaling all the time. This happens in both my local computer (BEAST v1.8.0) and the Cipres server (v1.7.5). I wonder if the tons of underflows warns me to reduce steps? Will less steps significantly impair the accuracy of the analysis?
Besides, I have got a bunch of test results with 50M + 500K x 50 steps and compared them to my previous 25M + 100K x 100 results. The absolute values appear to be smaller (more negative). I have not yet got several replicates per scenario so I cannot tell how stable the result is by far.
Best,
Qiyun
Thanks for the explanation. Yes I used dynamic scaling all the time. This happens in both my local computer (BEAST v1.8.0) and the Cipres server (v1.7.5). I wonder if the tons of underflows warns me to reduce steps? Will less steps significantly impair the accuracy of the analysis?
Besides, I have got a bunch of test results with 50M + 500K x 50 steps and compared them to my previous 25M + 100K x 100 results. The absolute values appear to be smaller (more negative). I have not yet got several replicates per scenario so I cannot tell how stable the result is by far.
Best,
Qiyun
Guy Baele
Jan 22, 2014, 4:38:25 AM1/22/14
to beast...@googlegroups.com
Hi Qiyun,
Op dinsdag 21 januari 2014 22:42:32 UTC+1 schreef Qiyun Zhu:
It's possible that less steps will prevent your marginal likelihood estimation process from reaching convergence.
Those underflows occur when moving closer to the prior, i.e. when less and less data is still "available" for the likelihood calculations.
As you state, a more computationally demanding analysis yields values that are different from the ones you had before.
This means that your original results were probably not correct, i.e. had failed to reach convergence.
This means that your original results were probably not correct, i.e. had failed to reach convergence.
Further increasing the settings might change your results even more ...
Best regards,
Guy
Op dinsdag 21 januari 2014 22:42:32 UTC+1 schreef Qiyun Zhu:
Qiyun Zhu
Jan 23, 2014, 3:35:34 AM1/23/14
to beast...@googlegroups.com
Hello Guy,
I see. Thanks! The length of chain I ran is almost the limitation of our computation resource. So it seems that I have to live with it. I have completed three replicates for one analysis. The results are:
rep AICM SS
1 414654.087 -209395.73
2 414691.238 -209397.46
3 414574.002 -209402.02
The std.dev for AICM is 59.9 and that for SS is only 3.25. I guess this means that SS is indeed more accurate than AICM. Yeh~
Best,
Qiyun
I see. Thanks! The length of chain I ran is almost the limitation of our computation resource. So it seems that I have to live with it. I have completed three replicates for one analysis. The results are:
rep AICM SS
1 414654.087 -209395.73
2 414691.238 -209397.46
3 414574.002 -209402.02
The std.dev for AICM is 59.9 and that for SS is only 3.25. I guess this means that SS is indeed more accurate than AICM. Yeh~
Best,
Qiyun
Guy Baele
Jan 23, 2014, 4:20:17 AM1/23/14
to beast...@googlegroups.com
Hi Qiyun,
Op donderdag 23 januari 2014 09:35:34 UTC+1 schreef Qiyun Zhu:
No, that's not what it means at all !
It simply means that the variation between runs on the AICM is higher than on SS.
But if the variation occurs around the wrong value, even though that variation can be smaller, it's still wrong.
Besides, the AICM is not a marginal likelihood estimator, so it's useless comparing the two using a standard deviation.
Besides, the AICM is not a marginal likelihood estimator, so it's useless comparing the two using a standard deviation.
You need to check whether the SS estimate is consistent/stable when applying additional computational resources to estimate it.
E-mail me if you'd like to discuss your results in more detail.
Best regards,
Guy
Guy
Op donderdag 23 januari 2014 09:35:34 UTC+1 schreef Qiyun Zhu:
Qiyun Zhu
Jan 23, 2014, 11:58:20 AM1/23/14
to beast...@googlegroups.com
Hello Guy,
Oh given that I will try to do a more robust computation and see if results get stable. Maybe do 1000K x 100 steps and compare to 500K x 50 steps.Qiyun
--
You received this message because you are subscribed to a topic in the Google Groups "beast-users" group.
To unsubscribe from this topic, visit https://groups.google.com/d/topic/beast-users/vaR6fxQUXVQ/unsubscribe.
To unsubscribe from this group and all its topics, send an email to beast-users...@googlegroups.com.
To post to this group, send email to beast...@googlegroups.com.
Visit this group at http://groups.google.com/group/beast-users.
For more options, visit https://groups.google.com/groups/opt_out.
Guy Baele
Jan 23, 2014, 3:47:23 PM1/23/14
to beast...@googlegroups.com
Hi Qiyun,
Op donderdag 23 januari 2014 17:58:20 UTC+1 schreef Qiyun Zhu:
Yes, you could remove some samples from the end of each power posterior (and hence not only at the end of the file).
If you'd only cut from the end of the file, you'll only remove samples from the prior.
Best regards,
Guy
Op donderdag 23 januari 2014 17:58:20 UTC+1 schreef Qiyun Zhu:
Message has been deleted
Qiyun Zhu
Jan 23, 2014, 6:32:24 PM1/23/14
to beast...@googlegroups.com
Hello Guy,
I assume the "each power posterior" stands for the same "pathLikelihood.theta" value in the "mle.log" file, right?
So
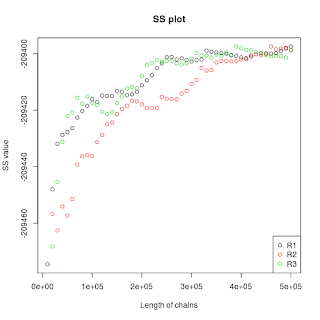
I cut of some samples from the end of each power posterior and had
BEAST calculate the SS. Then I plot the SS values of all three
replicates against chain length. Here is the plot:
Do you think it is already convergent. If not, how many more chain lengths (current: 500,000) and/or number of steps (current: 50) should I try? Because I have encounterred lots of underflows in the end of the 50 steps I'm fearing that adding more steps will be technically impossible.
I also attached the script I wrote to do this batch analysis, in case anyone wants to try. Usage: perl plotSS.pl xxx.mle.log
Best,
Qiyun
Guy Baele
Jan 24, 2014, 5:03:15 AM1/24/14
to beast...@googlegroups.com
Hi Qiyun,
Op vrijdag 24 januari 2014 00:32:24 UTC+1 schreef Qiyun Zhu:
Yes, that's correct.
What I find strange about your figure is that, in my experience, increased computational settings lead to lower log marginal likelihoods ...
A lot of people spend 100 million iterations or more on their parameter inference, so given that PS/SS have a reputation of being computationally challenging, at least the same number of iterations can be expected in total to perform the ML estimations.
And I would want the curves to flatten out to indicate convergence, which in my opinion is not yet really the case here.
We're currently testing if splitting up the PS/SS calculations is a good approach (with respect to convergence issues) to tackling these computational issues.
Best regards,
Guy
Guy
Op vrijdag 24 januari 2014 00:32:24 UTC+1 schreef Qiyun Zhu:
Qiyun Zhu
Jan 26, 2014, 11:57:58 AM1/26/14
to beast...@googlegroups.com
Hello Guy,
Thanks! i raised the iterations from 500K to 1M and wait to see if the curve gets convergent.
Best,Thanks! i raised the iterations from 500K to 1M and wait to see if the curve gets convergent.
Qiyun
Qiyun Zhu
Mar 4, 2014, 12:35:59 PM3/4/14
to beast...@googlegroups.com
Hello Guy,
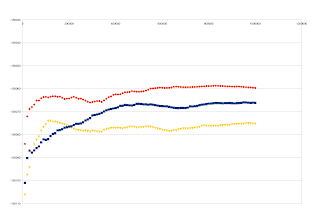
I'm back to this thread again with my new results, which is a maximum of our computational resource: 100M (original MCMC) + 1M x100 steps (PS/SS MCMC). It seems that the curves converge better but still vary among replicates. Below are the SS vs. #Generation curves of three different topological hypotheses, each having three replicates.
I took one of these hypotheses and combine the three replicates. It looks like this:
I found that the three replicates of the original MCMC seem to converge well, like this:
How do you think of these results? Are they good enough to be final results? Should I report the merge of three replicates or should I only report the most stable one of them? I really appreciate if you can share your opinions on this. Thanks!
Best regards,
Qiyun
Guy Baele
Mar 4, 2014, 3:04:41 PM3/4/14
to beast...@googlegroups.com
Hi Qiyun,
Op dinsdag 4 maart 2014 18:35:59 UTC+1 schreef Qiyun Zhu:
I understand that the posterior traces (and hence the AICM, HME, sHME) seem to be more stable. Unfortunately, more stable does not mean more accurate.
And I think that making claims of stability based on the graphs you show is very dangerous and I would hence not do so.
Your strategy of running the same XML is a definite plus, since that's also advocated for standard MCMC analyses.
The main problem that I have with your results (I have had this as well) is that different runs produce different outcomes, so no, I don't think this will support the claim you're trying to make concerning the different topological hypotheses.
Looking more closely at the results, you can see why. Between runs there is a difference of up to 60 log units between the runs.
And it would seem that the true difference in log marginal likelihood between the different hypothesis is smaller than that, preventing you from drawing a strong conclusion.
In other words, your log marginal likelihood estimates have unfortunately not converged yet.
How long did these runs take to finish?
E-mail me (I'm away until the end of the week though) with more details and we can see if we can work something out to reach convergence.
Best regards,
Guy
Op dinsdag 4 maart 2014 18:35:59 UTC+1 schreef Qiyun Zhu:
Reply all
Reply to author
Forward
0 new messages