GFF3 error :incorrect gff
279 views
Skip to first unread message
Juliana Chu
Feb 4, 2021, 1:30:59 AM2/4/21
to majiq_voila
Hello, there!
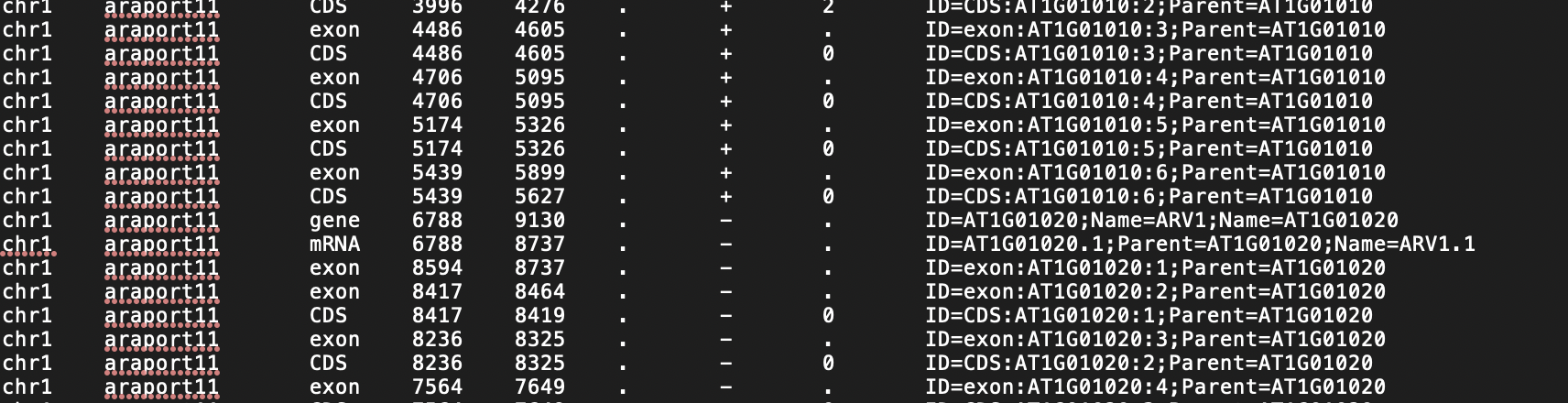
For the gff3 file, I have converted the gtf to gff3 file, with a script I wrote. However, it gives me errors for every line: WARNING - Error, incorrect gff. exon doesn't have valid mRNA b'CDS:AT1G01010:1', and with value error at the end :ValueError: not enough values to unpack (expected 2, got 1)
This is an example of my gff3 file:

I think it is mostly same format as the example give in workshop.
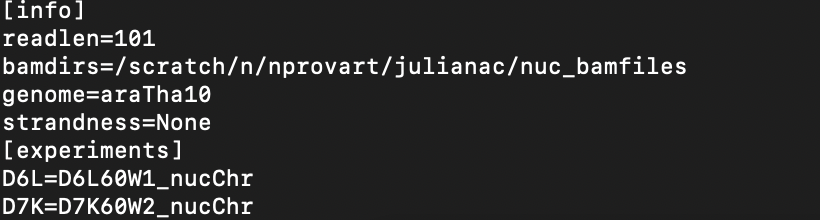
and this is my configuration file:

Is it wrong with the genome in the configuration file? or is my gff3 file still not complete? Please help me!
Thank you!
-Juliana
Paul Jewell
Feb 10, 2021, 4:11:39 PM2/10/21
to majiq_voila
Hi Juliana,
I'm thinking this is the same issue as the thread at https://groups.google.com/g/majiq_voila/c/5XIEfD-Zo9I . Is that correct? If so, I will close this one.
Thanks,
-Paul
Reply all
Reply to author
Forward
0 new messages
