Question about the `total_flux` dataset in the direct.h5 file.
143 views
Skip to first unread message
jintu zhang
Dec 3, 2022, 7:40:39 AM12/3/22
to westpa-users
Dear all,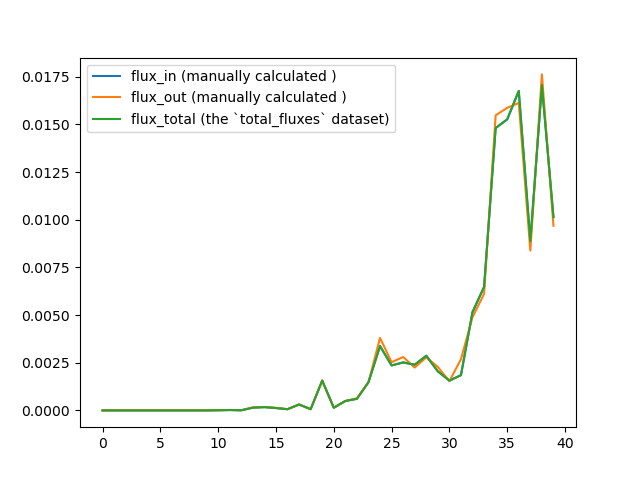
I would like to seek help with the `total_flux` dataset in the direct.h5 file.
From the wiki page, I know that this dataset represents the total flux into each macrostate within a WE iteration. So for example, with a four-state system, the total flux into state 2 would equal J_02 + J_12 + J_32. Thus I checked this relationship with the provided west.h5 file of the p53 example on GitHub. I slightly modified the west.cfg file to split the pcoord space into 4 states. After running the `w_ipa -ao` command, I summed up the fluxes into state 2 with a simple python script (the one_state_flux.py file is attached to this email). But it seems that the result given by the `total_flux` dataset is not consistent with the manually calculated results:

So I wonder if I have made any mistakes. Thanks in advance for the help!
Jintu Zhang
PhD student at Zhejiang University
Anthony Bogetti
Dec 3, 2022, 10:57:53 AM12/3/22
to westpa...@googlegroups.com
Hello Jintu,
Thank you for your question; we are happy to help you solve it. I ran a w_ipa analysis on the same h5 file with the analysis scheme from the attached west.cfg file. From my analysis, I found that indeed the conditional flux from 0 -> 2 + 1 -> 2 + 3 -> 2 is equal to the total flux into 2 at every time point during the simulation (I did a check of `if flux_in[i] != flux_total[i,2]:`. You are correct in thinking that these should be equal, and I think they are in your case. From the plot you attached, the green and blue lines should be overlapping, indicating the results of both methods of totaling the flux are the same. I attached my version where the flux_in and flux_total datasets are overlapping.
jintu zhang
Dec 3, 2022, 1:02:23 PM12/3/22
to westpa-users
Dear Anthony,
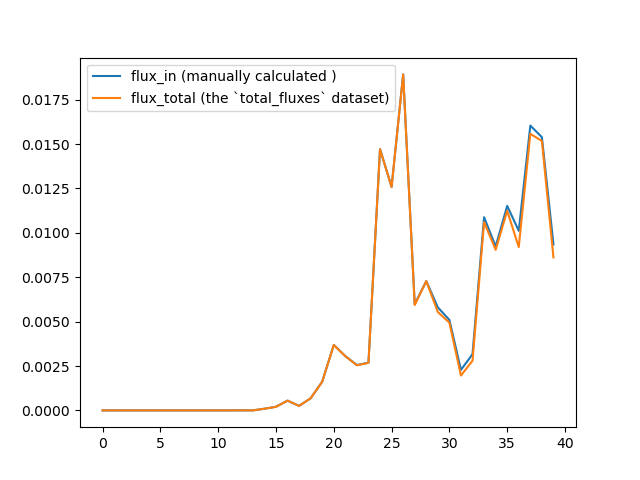
Thank you so much for the speedy reply! I found that I made a silly mistake in my last post: just as you pointed out, the manually calculated fluxes and the WESTPA calculated fluxes are indeed the same. But when I tried to change the binning scheme, these results seems to disagree again:

I attached the updated west.cfg to the email. Could you please revisit the question? Thanks in advance!
Jintu Zhang
Anthony Bogetti
Dec 7, 2022, 11:17:51 AM12/7/22
to westpa...@googlegroups.com
Hello Jintu,
I was able to reproduce this discrepancy using the same simulation data and don’t have an immediate answer for why the fluxes disagree. However, I do have a suggestion. Try breaking down the different state-to-state transitions into individual steps as schemes in your west.cfg’s analysis section. For instance, instead of looking at 0 to 1, 0 to 2, 0 to 3, 1 to 2 etc. all within the same scheme, do something like the following:
0_to_1:
enabled: True
bins:
- type: RectilinearBinMapper
boundaries:
- [0.0, 10.0, 'inf']
- [0.0, 10.0, 'inf']
states:
- label: s0
coords:
- [1.0, 1.0]
- label: s1
coords:
- [11.0, 1.0]
0_to_2:
enabled: True
bins:
- type: RectilinearBinMapper
boundaries:
- [0.0, 10.0, 'inf']
- [0.0, 10.0, 'inf']
states:
- label: s0
coords:
- [1.0, 1.0]
- label: s2
coords:
- [1.0, 11.0]
And so forth for all of the transitions of interest.
It may be the case that within a given iteration, transitions that pass through multiple states either count or don’t count towards the conditional flux. The above suggestion where everything is broken down into transitions between two states might be cleaner and give more consistent results. Try it out and let us know!
Also, generally speaking, we usually only look at transitions between two states at a time in the west.cfg’s analysis section when we do analysis, not only for this reason, but just because it makes data selection from the resulting direct.h5 files less error-prone.
Best,
Anthony
On Dec 3, 2022, at 1:02 PM, jintu zhang <zhan...@gmail.com> wrote:
Dear Anthony,Thank you so much for the speedy reply! I found that I made a silly mistake in my last post: just as you pointed out, the manually calculated fluxes and the WESTPA calculated fluxes are indeed the same. But when I tried to change the binning scheme, these results seems to disagree again:
<flux.png>I attached the updated west.cfg to the email. Could you please revisit the question? Thanks in advance!Jintu Zhang在2022年12月3日星期六 UTC+8 23:57:53<Anthony Bogetti> 写道:Hello Jintu,Thank you for your question; we are happy to help you solve it. I ran a w_ipa analysis on the same h5 file with the analysis scheme from the attached west.cfg file. From my analysis, I found that indeed the conditional flux from 0 -> 2 + 1 -> 2 + 3 -> 2 is equal to the total flux into 2 at every time point during the simulation (I did a check of `if flux_in[i] != flux_total[i,2]:`. You are correct in thinking that these should be equal, and I think they are in your case. From the plot you attached, the green and blue lines should be overlapping, indicating the results of both methods of totaling the flux are the same. I attached my version where the flux_in and flux_total datasets are overlapping.
--
You received this message because you are subscribed to the Google Groups "westpa-users" group.
To unsubscribe from this group and stop receiving emails from it, send an email to westpa-users...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/westpa-users/dbee454d-5543-4328-827e-c9a6efe12a38n%40googlegroups.com.
<west.cfg><flux.png>
Jintu Zhang
Dec 8, 2022, 4:28:44 AM12/8/22
to westpa-users
Dear Anthony,
Firstly, I am sorry about accidentally sending you two extra emails with the same content, I was not meant to do that. For the continuity of this thread, I decided to post that email once again here. I am sorry for the inconvenience, and hope you could understand :)
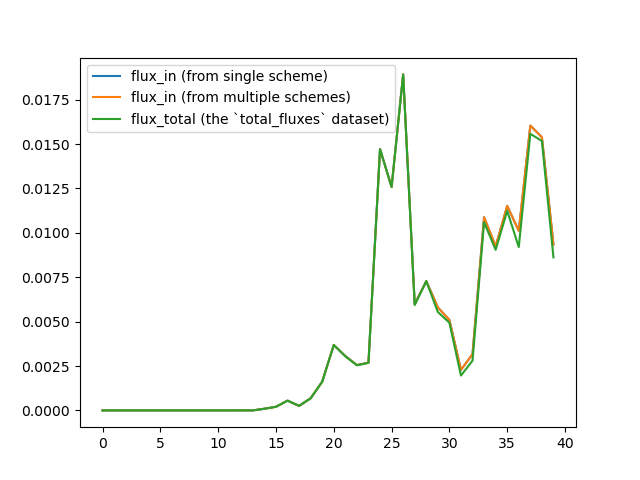
I have tested splitting the different state-to-state transitions into analysis schemes, but I got identical results:

The reason why I would like to obtain all the state-to-state fluxes is: I want to build a Markov State Model (not a haMSM but a relatively 'trivial' one with the WE bins as the MSM states) with these fluxes. So do you have any advice about such tasks? I really appreciate your help and support!
Jintu Zhang
Leung, Jeremy
Dec 16, 2022, 4:53:39 PM12/16/22
to westpa...@googlegroups.com
Hi Jintu,
Regarding the "rough" MSM construction, I would suggest using msm_we and just set 1 cluster per bin. The readthedocs there should be quite comprehensive. Here's the procedure if you decide to do it manually without
msm_we.
I would suggest just grabbing the state labels from assign.h5 and construct the dataset by hand. That way you have a better control about the lag time value you choose.
1. Create a n x n "count matrix" and zero it out. For a chosen lag time (example with your WE Tau here), loop through each segment and look at the last frame and its parent's state label (last frame of its parent). Add the weight of
that frame to the corresponding transition element in the "count matrix' .
2. Row normalize such that probability of each state sums up to 1. This will be your transition matrix.
3. Put the transition matrix into the chosen MSM program of your choice (PyEmma etc.) and continue. You have now interjected WE data into the program and can follow through.
Hope that helps. We're still looking into the w_direct discrepancies but we think it's related to the state definitions.
Regards,
Jeremy Leung
--
Jeremy M. G. Leung
PhD Candidate, Chemistry
Graduate Student Researcher, Chemistry (Chong Lab)
University of Pittsburgh | 219 Parkman Avenue, Pittsburgh, PA 15260
jml...@pitt.edu | [He, Him, His]
Jeremy M. G. Leung
PhD Candidate, Chemistry
Graduate Student Researcher, Chemistry (Chong Lab)
University of Pittsburgh | 219 Parkman Avenue, Pittsburgh, PA 15260
jml...@pitt.edu | [He, Him, His]
From: westpa...@googlegroups.com <westpa...@googlegroups.com> on behalf of Jintu Zhang <zhan...@gmail.com>
Sent: Thursday, December 8, 2022 4:28:44 AM
To: westpa-users <westpa...@googlegroups.com>
Subject: Re: [westpa-users] Question about the `total_flux` dataset in the direct.h5 file.
Sent: Thursday, December 8, 2022 4:28:44 AM
To: westpa-users <westpa...@googlegroups.com>
Subject: Re: [westpa-users] Question about the `total_flux` dataset in the direct.h5 file.
Dear Anthony,
Firstly, I am sorry about accidentally sending you two extra emails with the same content, I was not meant to do that. For the continuity of this thread, I decided to post that email once again here. I am sorry for the inconvenience, and hope you could
understand :)
I have tested splitting the different state-to-state transitions into analysis schemes, but I got identical results:
To view this discussion on the web visit
https://groups.google.com/d/msgid/westpa-users/68f5991a-d767-41ba-861e-0c07d157bf46n%40googlegroups.com.
Jintu Zhang
Dec 20, 2022, 5:47:18 AM12/20/22
to westpa-users
Dear Jeremy,
Thank you very much for the answers! These are really helpful.
Jintu Zhang
Reply all
Reply to author
Forward
0 new messages
