Using pcount() to correct for imperfect detection for two seasons of data: looking for procedure input and feedback on resulting abundance inflation issues
757 views
Skip to first unread message
Tim Duclos
Jan 5, 2016, 4:53:10 PM1/5/16
to unmarked
Greetings Group,
![]()
I would like to gather feedback on my N-Mix modeling approach. Any input is very appreciated!
I have 3 replicate point counts of avian species from 100 sample sites over two seasons, respectfully. Additionally, I repeatedly sampled 50 new sites in my second season. Thus, overall I have 3 replicate counts at 100 sites in 2014 and 3 replicate counts at 150 sites in 2014 (i.e. 6 replicate counts at the same 100 sites over two years and 3 replicate counts at 50 sites in only one year). Make sense?
For each individual species, I would like to use pcount() to derive corrected abundance estimates at each site, for each year independently, utilizing only my detection covariates (obsCovs) and nothing else (i.e. ~1 on state side of the model).
Therefore my objective is to derive corrected abundance estimates for each site, each year, having taken into account imperfect detection. Seems simple enough, eh?
From what I gather, it seems that I want to stack my data like such:
obs 1, obs 2, obs 3,
point_1_year_14,
point_1_year_15,
point_2_year_14,
point_2_year_15,
...
Thus, my data setup is such:
zUMF<-with(bird.2,{
unmarkedFramePCount(
y = cbind(species_count.1,species_count.2,species_count.3), #define response variable
siteCovs = data.frame(point_id_num), #enter point_id numeric dummy variable column as fixed effect (because cant handle character values)
obsCovs = list(year = cbind(year.1,year.2,year.3), #define detection covariates
date = cbind(date.1,date.2,date.3),
time = cbind(time.1,time.2,time.3),
observer = cbind(observer.1,observer.2,observer.3),
sky = cbind(sky.1,sky.2,sky.3),
wind = cbind(wind.1,wind.2,wind.3),
stream = cbind(stream.1,stream.2,stream.3),
temp_c = cbind(temp_c.1,temp_c.2,temp_c.3)))})
I have seven detection covariates that I have reason to believe either individually, or in combination, may affect detectability of these birds. Additionally, I have included year as a detection covariate in order to take into account the fact that my models include surveys at the same sites but from separate years (but debate whether this is logical given my assumption of site closure and whether or not year is redundant given date (which is julian date derived from: mm/dd/yy). I am using the dredge() function from the MuMin package to run models of all subsets of the seven detection covariates (with ~1 on state side of model), calculate AIC for each model, and thus identify the "best" model for each species containing only the significant detection covariates for the given species.
This is my global model presented to dredge():
global <- pcount(formula= ~ year + date + time + observer + sky + wind + stream + temp_c ~ 1 , data=zUMF, mixture = "ZIP")
I then confirm GOF of the resulting top model using parboot(). For all species ZIP fits these data well.
Next, I use the raneff() function to derive corrected abundance estimates for each site given the top model as identified in the dredge()
est<-data.frame(bup(ranef(top.mod), type = "mean"))
This results in a mean corrected abundance estimate for each site for each year across all surveys within year. This is exactly what I am looking for.
All this being said, I have two concerns which I would like feedback on from the group:
1) Overall, does this seem like a logical modeling procedure given my objective?
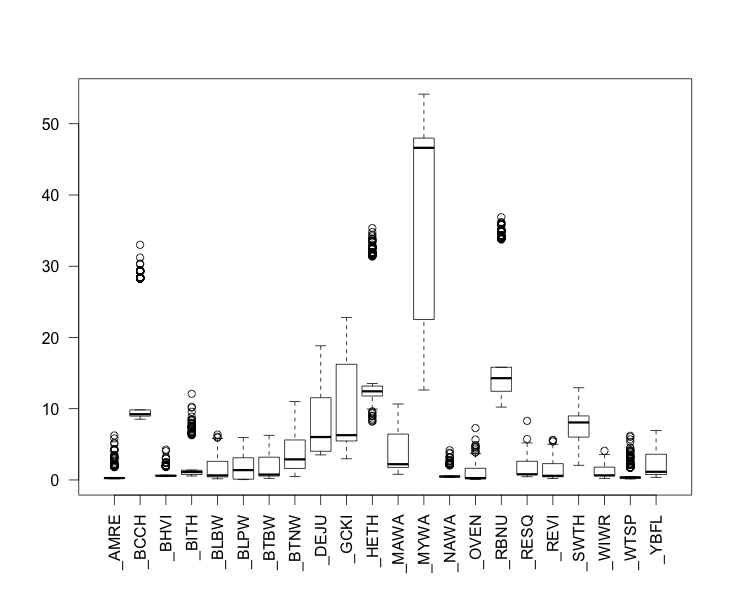
2) For some of these species, my resulting corrected abundance estimates are EXTREMELY inflated. For many others, there still seems to be undue variability and inflation in the abundance estimates. In the figure below you can see the results of my corrected abundance estimates for each species from the above procedure. Y-axis is corrected abundance estimate for each site. X-axis is species (AOU code).
From previous discussions on this forum I know that with counts that are very zero-inflated (as these are) you can get very low detection probabilities. Such low detection probabilities can lead to inflation of abundance estimates and overall instability in the abundance estimates- even when parboot() suggests that the GOF is good. Thus, as we all know, a well fit model can still be a bad model. From my investigation thus far, it seems that there is little to be done to mitigate such phenomenon. Is this a fair conclusion? Are there any options that exist to try and address such inflation, either pre or post modeling?
Overall, what steps can I take to troubleshoot these inflated results?
I hope I explained my procedure adequately. If not, I'll clarify as much as possible. Thank you in advance for any feedback you may have!
Cheers,
Tim
UMass Amherst
John Clare
Jan 5, 2016, 11:35:40 PM1/5/16
to unmarked
Hi Tim,
Just as an FYI, graphics aren't displaying (on my computer at least). It might also be helpful to paste tabular output and add a little additional context: there are a number of things that can make estimates wonky, and it might make distinguishing between issues related to sparse data vs. a mis-specified abundance model and/or mixing distribution or something else easier.
Cheers,
John
Just as an FYI, graphics aren't displaying (on my computer at least). It might also be helpful to paste tabular output and add a little additional context: there are a number of things that can make estimates wonky, and it might make distinguishing between issues related to sparse data vs. a mis-specified abundance model and/or mixing distribution or something else easier.
Cheers,
John
Message has been deleted
Tim Duclos
Jan 6, 2016, 1:49:24 PM1/6/16
to unmarked
Thank you for your response. Here is another attempt at attaching the figure of my corrected abundance estimates by species:
Also, I think I have found a potential cause for my inflated results. In the tabular output below for the BCCH dataframe you can see that the calculated mean number of observation per site is absolutely incorrect given the actual tabulation of observations. What gives?
zUMF<-with(bird.2,{
unmarkedFramePCount(
y = cbind(species_count.1,species_count.2,species_count.3), #define response variable
siteCovs = data.frame(point_id_num), #enter point_id numeric dummy variable column as fixed effect (because cant handle character values)
obsCovs = list(year = cbind(year.1,year.2,year.3),#define detection covariates
date = cbind(date.1,date.2,date.3),
time = cbind(time.1,time.2,time.3),
observer = cbind(observer.1,observer.2,observer.3),
sky = cbind(sky.1,sky.2,sky.3),
wind = cbind(wind.1,wind.2,wind.3),
stream = cbind(stream.1,stream.2,stream.3),
temp_c = cbind(temp_c.1,temp_c.2,temp_c.3)))})
summary(zUMF)
unmarkedFrame Object
250 sites
Maximum number of observations per site: 3
Mean number of observations per site: 3
Sites with at least one detection: 60
Tabulation of y observations:
0 1 2 3 <NA>
679 53 16 2 0
Site-level covariates:
point_id_num
Min. : 1.00
1st Qu.: 63.25
Median :125.50
Mean :125.50
3rd Qu.:187.75
Max. :250.00
Observation-level covariates:
year date time observer sky
Min. :14.0 Min. :16222 Min. :0.1868 Min. :1.000 Min. :0.000
1st Qu.:14.0 1st Qu.:16248 1st Qu.:0.2285 1st Qu.:1.000 1st Qu.:0.000
Median :15.0 Median :16591 Median :0.2743 Median :1.000 Median :1.000
Mean :14.6 Mean :16458 Mean :0.2751 Mean :1.567 Mean :1.211
3rd Qu.:15.0 3rd Qu.:16604 3rd Qu.:0.3222 3rd Qu.:2.000 3rd Qu.:2.000
Max. :15.0 Max. :16613 Max. :0.4028 Max. :3.000 Max. :5.000
wind stream temp_c
Min. :0.000 Min. :0.0000 Min. : 1.50
1st Qu.:0.000 1st Qu.:0.0000 1st Qu.: 9.00
Median :1.000 Median :0.0000 Median :11.50
Mean :0.936 Mean :0.6147 Mean :11.34
3rd Qu.:1.000 3rd Qu.:1.0000 3rd Qu.:13.50
Max. :5.000 Max. :3.0000 Max. :22.00
John Clare
Jan 6, 2016, 9:00:53 PM1/6/16
to unmarked
Hi Tim,
That graphic worked, thanks. I think the 'mean # obs' actually refers to the number of visits or sampling occasions. I think that one thing that might be beneficial is to define year and observer as factors. At present, it looks like you are fitting a log-linear trend to both of these (won't matter for year because there are only 2 levels). Might think about coding wind, sky, and stream as factors as well, assuming these fall on some ordinal scale. Also, not quite sure what the time covariate is coding..., and the date covariate is so large that it may be causing fitting issues. Presumably, you could divide the julian day by 10 or 100 or something.
Cheers,
John
Tim Duclos
Jan 11, 2016, 6:42:56 PM1/11/16
to unmarked
Hi John,
Thank you very much for your feedback. Indeed most of my covariates are on the ordinal scale, and I could see why they should be considered as factors, as you suggest. I tried this- but the models will not converge when the covariates are coded as factors (???)- I get the "Hessian is singular- try fewer covariates" error. I guess this error is thrown when one or more parameters cannot be estimated. Anyhow, I'm going to keep trying and will post my progress.
Also, I appreciate the idea of rescaling julian date (by dividing by 1000)- this will likely make the models run faster, at the very least.
Best,
Tim
John Clare
Jan 12, 2016, 4:02:02 PM1/12/16
to unmarked
Hi Tim,
Good luck. Just to follow-up quickly regarding factors vs. covariates: for something like sky, if 1 ~0-20%, 2 ~20-40%,...5~80-100% cloud or whatever, you might be able to get away with treating it as continuous. And re. any continuous covariate, e.g., temp, date, time, simply scaling may be sufficient for letting the algorithm find an optima, or scaling and centering (i.e., converting to Z-scores) might be necessary. I think most people would recommend scaling and centering as the safest bet, although I've never found an estimate that differed beyond rounding error once both were back-transformed to the same scale.
But a disclaimer for consideration before spending a bunch of time doing things that somebody online recommends: these suggestions are more aimed at assisting the fitting algorithm and treating sources of variance sensibly, and may very well have little effect regarding correcting the inflated abundance issue. Taking a look at some of the primary literature that discusses potential sources of bias in N-mixture models (very low p, violations of closure/independence, etc.) might help narrow the issue. One thing to keep in mind is that a detection-corrected count derived using an N-mixture model is not quite the same as a detection corrected count derived from a Huggins model or something: abundance is still a part of the likelihood, and structuring the model such that variation is predominantly or exclusively within detection can result in squeezing the abundance distribution into a box where it doesn't quite fit. If there is true variation in abundance, your finite-sample estimates of N should be more accurate if the source of that variance is explicitly modeled.
Tim Duclos
Jan 18, 2016, 4:29:24 PM1/18/16
to unmarked
Hi John et al.
So I have landed on a solid hypothesis explaining my low detection probability and the resulting inflated abundance estimates from pcount(). Looking closely at the detection history for individual species as each site, there is a lot of heterogeneity in the count values between repeat visits. I.e. for example it was not uncommon to have, say, 3 individuals counted at a site on the first visit, none on the second visit, and say, 1 on the third visit. This is just an example- but you get the point. Basically, the more heterogeneous/variable your detection history at a site, the lower the detection probability will likely be. I have a fair amount of such variability and thus a fair amount of inflation caused by very low detection probability.
Barring surveyor error, I feel like such variability in detection history could likely be indicative of an open population rather than a closed one (as I have thus far assumed). First, it does seem reasonable to me to accept that a 50m radius sample plot could occur at the intersection of the fringe territories of multiple individuals of the same species. Second, based upon my personal experience (and well documented in the lit) it is known that birds sing in bouts- often stimulated by the singing of other individuals. Thus, I could see where in one survey you could detect multiple individuals singing on their respective territories as they intersect on the survey plot, perhaps in response to one another- but then in subsequent surveys you may not detect these individuals because they are potentially off the plot on another part of their territory (but still in the general vicinity) or, undeniably, perhaps on the plot but not vocalizing or otherwise undetectable. This seems logical and a good explanation for variability in detection history.
Thus, I am wondering how I should test for openness in the population.
I should also mention that I had originally subdivided my 10-minute point counts into four 2.5m sub intervals during which time I kept track of each individual detected as well as the interval in which I originally detected each individual. This is the so-called "time-of-detection" sampling method. Therefore, I am wondering if, and how, I could/should use my sub-divided detection history to account for variability in singing rates and thus take into account singing rate when calculating detection probability. Additionally, it seems that I should also use pcountOpen() to allow for immigration and emigration between surveys. Is there a method in Unmarked to both utilize detection history at the sub-interval level (account for singing rates) as well as allow for openness between surveys?
What do y'all think?
Any input is welcomed and helpful!
-Tim
John Clare
Jan 18, 2016, 5:20:06 PM1/18/16
to unmarked
Hi Tim,
Might try looking at unmarked's gmultmix and the paper describing it (Chandler et al. 2011). I believe that Dail and Madsen (2011) describe some ways to formally test for closure (maybe this was as straight-forward as a L-R test). Primary difference between gmultmix and pcountOpen is whether you prefer dynamics modeled explicitly (in terms of #arrivals and %remaining, which may be analogous to recruitment and apparent survival), or whether the dynamics are related to temporary random emigration (i.e., not always availabile). To me, gmultmix seems more in line with what you are interested in, but perhaps others feel differently?
John
Dan Linden
Jan 19, 2016, 12:33:46 PM1/19/16
to unmarked
Yes, I'd say that gmultmix with a removal design makes the most sense here. Given that the replicate surveys are within a time frame that could reasonably be considered closed to population dynamics, the true mechanism driving the observations is likely temporary emigration (especially with a 50-m plot size).
Tim, you can look to the gmultmix help file for an example, but you'll want your data formatted such that each of the 2.5 min intervals indicates the number of newly detected individuals. For you, the number of primary periods is the number of replicates (T=3) and the number of secondary periods is the number of 2.5 min intervals (J=4).
Tim, you can look to the gmultmix help file for an example, but you'll want your data formatted such that each of the 2.5 min intervals indicates the number of newly detected individuals. For you, the number of primary periods is the number of replicates (T=3) and the number of secondary periods is the number of 2.5 min intervals (J=4).
Tim Duclos
Jan 21, 2016, 4:06:13 PM1/21/16
to unmarked
John and Dan, Thank you so much for your reply and much valued input! Dan, given that you seem to have worked with the gmultmix model before, I wonder if you (or anybody else on here that has used this model before) could help me set up my model. Here is what I've got thus far:
As a reminder, I am looking to derive corrected abundance estimates for species observed on each of 150 plots over two consecutive years. Each plot was sampled via standard point count protocol 3 times each season, and each of these repeat survey were divided into 4 intervals of equal time, for which I have counts of newly detected individuals by species (i.e “time-of-detection” sample method). Furthermore I recorded data on 7 different detection covariates at the time of survey. Given heterogeneity in count values across repeat visits (i.e. large variability in detection history) I would like to take into account temporary emigration between surveys when deriving these corrected abundance estimates. To this end, it seems that the gmultmix() model within the Unmarked package in R is what I want to work with. FYI i have read the Unmarked help file as well as the paper describing this model titled “Inference about density and temporary emigration in unmarked populations” (Chandler et al. 2011).
Note: I would like to also assume closure within season, so I want to derive abundance estimates for each sample site individually with respect to year- thus I have simply stacked the data and recoded the site ID to include year so as to be treated as separate sites between years.
Yet I am a bit confused as how to present my data to the gmultmix model. First, I want to ensure that I have formatted my data correctly within the unmarkedFrameGMM. Here is my data frame that I am working to format into the unmarkedFrameGMM:
Note that I have already formatted the data into wide format such that each row represents a unique sample site and the columns are formatted such that the first number following the dot represents data from a particular replicate survey and the last number represents data from an interval within the survey (ex: .1_1 means first rep and first interval within that rep). Here you can see that year, date, time, observer, sky, wind, stream, and temp_c are my detection covariates (FYI I am using characters to constrain the non-continuous covariate values as factors). Note that these covariates do not vary across intervals within a replicate survey but only between replicate surveys. Species_count varies at the interval level and represents the number of new individuals detected within one of four intervals within a given replicate survey.
> str(bird.2)
'data.frame': 250 obs. of 110 variables:
$ point_id : Factor w/ 150 levels "ADAM01","ADAM02",..: 1 1 2 2 3 3 4 4 5 5 ...
$ point_id_year : chr "ADAM01_A" "ADAM01_B" "ADAM02_A" "ADAM02_B" ...
$ year.1_1 : chr "A" "B" "A" "B" ...
$ date.1_1 : num 16236 16593 16236 16593 16236 ...
$ time.1_1 : num 0.283 0.295 0.308 0.322 0.338 ...
$ observer.1_1 : Factor w/ 3 levels "ML","SD","TD": 3 3 3 3 3 3 3 3 3 3 ...
$ sky.1_1 : chr "E" "A" "E" "A" ...
$ wind.1_1 : chr "C" "A" "B" "B" ...
$ stream.1_1 : chr "A" "B" "A" "A" ...
$ temp_c.1_1 : num 10 6.5 10 6 9 5.5 7 7.5 14 3.5 ...
$ species_count.1_1: num 0 1 1 0 0 1 0 0 0 0 ...
$ year.1_2 : chr "A" "B" "A" "B" ...
$ date.1_2 : num 16236 16593 16236 16593 16236 ...
$ time.1_2 : num 0.283 0.295 0.308 0.322 0.338 ...
$ observer.1_2 : Factor w/ 3 levels "ML","SD","TD": 3 3 3 3 3 3 3 3 3 3 ...
$ sky.1_2 : chr "E" "A" "E" "A" ...
$ wind.1_2 : chr "C" "A" "B" "B" ...
$ stream.1_2 : chr "A" "B" "A" "A" ...
$ temp_c.1_2 : num 10 6.5 10 6 9 5.5 7 7.5 14 3.5 ...
$ species_count.1_2: num 0 0 0 0 0 0 0 0 0 0 ...
$ year.1_3 : chr "A" "B" "A" "B" ...
$ date.1_3 : num 16236 16593 16236 16593 16236 ...
$ time.1_3 : num 0.283 0.295 0.308 0.322 0.338 ...
$ observer.1_3 : Factor w/ 3 levels "ML","SD","TD": 3 3 3 3 3 3 3 3 3 3 ...
$ sky.1_3 : chr "E" "A" "E" "A" ...
$ wind.1_3 : chr "C" "A" "B" "B" ...
$ stream.1_3 : chr "A" "B" "A" "A" ...
$ temp_c.1_3 : num 10 6.5 10 6 9 5.5 7 7.5 14 3.5 ...
$ species_count.1_3: num 0 0 1 0 0 0 0 0 0 0 ...
$ year.1_4 : chr "A" "B" "A" "B" ...
$ date.1_4 : num 16236 16593 16236 16593 16236 ...
$ time.1_4 : num 0.283 0.295 0.308 0.322 0.338 ...
$ observer.1_4 : Factor w/ 3 levels "ML","SD","TD": 3 3 3 3 3 3 3 3 3 3 ...
$ sky.1_4 : chr "E" "A" "E" "A" ...
$ wind.1_4 : chr "C" "A" "B" "B" ...
$ stream.1_4 : chr "A" "B" "A" "A" ...
$ temp_c.1_4 : num 10 6.5 10 6 9 5.5 7 7.5 14 3.5 ...
$ species_count.1_4: num 0 0 0 0 0 0 0 0 0 0 ...
$ year.2_1 : chr "A" "B" "A" "B" ...
$ date.2_1 : num 16249 16604 16249 16604 16249 ...
$ time.2_1 : num 0.265 0.275 0.238 0.249 0.214 ...
$ observer.2_1 : Factor w/ 3 levels "ML","SD","TD": 3 2 3 2 3 2 3 2 3 2 ...
$ sky.2_1 : chr "A" "B" "A" "C" ...
$ wind.2_1 : chr "B" "A" "B" "B" ...
$ stream.2_1 : chr "A" "A" "A" "B" ...
$ temp_c.2_1 : num 13 10.5 12 10 11 9 9 8 17 13.5 ...
$ species_count.2_1: num 0 0 1 0 1 0 0 0 0 0 ...
$ year.2_2 : chr "A" "B" "A" "B" ...
$ date.2_2 : num 16249 16604 16249 16604 16249 ...
$ time.2_2 : num 0.265 0.275 0.238 0.249 0.214 ...
$ observer.2_2 : Factor w/ 3 levels "ML","SD","TD": 3 2 3 2 3 2 3 2 3 2 ...
$ sky.2_2 : chr "A" "B" "A" "C" ...
$ wind.2_2 : chr "B" "A" "B" "B" ...
$ stream.2_2 : chr "A" "A" "A" "B" ...
$ temp_c.2_2 : num 13 10.5 12 10 11 9 9 8 17 13.5 ...
$ species_count.2_2: num 0 0 0 0 0 0 0 0 0 0 ...
$ year.2_3 : chr "A" "B" "A" "B" ...
$ date.2_3 : num 16249 16604 16249 16604 16249 ...
$ time.2_3 : num 0.265 0.275 0.238 0.249 0.214 ...
$ observer.2_3 : Factor w/ 3 levels "ML","SD","TD": 3 2 3 2 3 2 3 2 3 2 ...
$ sky.2_3 : chr "A" "B" "A" "C" ...
$ wind.2_3 : chr "B" "A" "B" "B" ...
$ stream.2_3 : chr "A" "A" "A" "B" ...
$ temp_c.2_3 : num 13 10.5 12 10 11 9 9 8 17 13.5 ...
$ species_count.2_3: num 0 0 0 0 0 0 0 0 0 0 ...
$ year.2_4 : chr "A" "B" "A" "B" ...
$ date.2_4 : num 16249 16604 16249 16604 16249 ...
$ time.2_4 : num 0.265 0.275 0.238 0.249 0.214 ...
$ observer.2_4 : Factor w/ 3 levels "ML","SD","TD": 3 2 3 2 3 2 3 2 3 2 ...
$ sky.2_4 : chr "A" "B" "A" "C" ...
$ wind.2_4 : chr "B" "A" "B" "B" ...
$ stream.2_4 : chr "A" "A" "A" "B" ...
$ temp_c.2_4 : num 13 10.5 12 10 11 9 9 8 17 13.5 ...
$ species_count.2_4: num 0 0 0 0 0 0 0 0 0 0 ...
$ year.3_1 : chr "A" "B" "A" "B" ...
$ date.3_1 : num 16262 16613 16262 16613 16262 ...
$ time.3_1 : num 0.284 0.29 0.31 0.319 0.334 ...
$ observer.3_1 : Factor w/ 3 levels "ML","SD","TD": 3 3 3 3 3 3 3 3 3 3 ...
$ sky.3_1 : chr "B" "A" "A" "A" ...
$ wind.3_1 : chr "A" "A" "A" "A" ...
$ stream.3_1 : chr "A" "A" "A" "A" ...
$ temp_c.3_1 : num 9 9.5 7 9.5 7 9.5 10 12 7 8.5 ...
$ species_count.3_1: num 0 1 0 0 0 0 0 0 0 0 ...
$ year.3_2 : chr "A" "B" "A" "B" ...
$ date.3_2 : num 16262 16613 16262 16613 16262 ...
$ time.3_2 : num 0.284 0.29 0.31 0.319 0.334 ...
$ observer.3_2 : Factor w/ 3 levels "ML","SD","TD": 3 3 3 3 3 3 3 3 3 3 ...
$ sky.3_2 : chr "B" "A" "A" "A" ...
$ wind.3_2 : chr "A" "A" "A" "A" ...
$ stream.3_2 : chr "A" "A" "A" "A" ...
$ temp_c.3_2 : num 9 9.5 7 9.5 7 9.5 10 12 7 8.5 ...
$ species_count.3_2: num 0 0 1 0 0 0 0 0 0 0 ...
$ year.3_3 : chr "A" "B" "A" "B" ...
$ date.3_3 : num 16262 16613 16262 16613 16262 ...
$ time.3_3 : num 0.284 0.29 0.31 0.319 0.334 ...
$ observer.3_3 : Factor w/ 3 levels "ML","SD","TD": 3 3 3 3 3 3 3 3 3 3 ...
$ sky.3_3 : chr "B" "A" "A" "A" ...
$ wind.3_3 : chr "A" "A" "A" "A" ...
$ stream.3_3 : chr "A" "A" "A" "A" ...
[list output truncated]
- attr(*, "reshapeWide")=List of 5
..$ v.names: chr "year" "date" "time" "observer" ...
..$ timevar: chr "rep_interval"
..$ idvar : chr "point_id_year" "point_id"
..$ times : chr "1_1" "1_2" "1_3" "1_4" ...
..$ varying: chr [1:9, 1:12] "year.1_1" "date.1_1" "time.1_1" "observer.1_1" ...
Next, I set up the unmarkedFrameGMM as such:
names(bird.2)
zUMF<-with(bird.2,{
unmarkedFrameGMM(
y = cbind(species_count.1_1,species_count.1_2,species_count.1_3,species_count.1_4,species_count.2_1,species_count.2_2,species_count.2_3,species_count.2_4,species_count.3_1,species_count.3_2,species_count.3_3,species_count.3_4), #define response variable (y=site (row) by rep*interval (columns) matrix)
siteCovs = data.frame (point_id_year), #need to provide list of sample sites
yearlySiteCovs = list(year = cbind(year.1_1,year.2_1,year.3_1),#define detection covariates varying with rep (because detection covs are same across intervals, just need to supply cov data from the first interval of each rep (kind of messy, but it works)
date = cbind(date.1_1,date.2_1,date.3_1),
time = cbind(time.1_1,time.2_1,time.3_1),
observer = cbind(observer.1_1,observer.2_1,observer.3_1),
sky = cbind(sky.1_1,sky.2_1,sky.3_1),
wind = cbind(wind.1_1,wind.2_1,wind.3_1),
stream = cbind(stream.1_1,stream.2_1,stream.3_1),
temp_c = cbind(temp_c.1_1,temp_c.2_1,temp_c.3_1)),
numPrimary=3, type="removal")})
summary(zUMF)
unmarkedFrame Object
250 sites
Maximum number of observations per site: 12
Mean number of observations per site: 12
Number of primary survey periods: 3
Number of secondary survey periods: 4
Sites with at least one detection: 128
Tabulation of y observations:
0 1 2 3 <NA>
2761 197 37 5 0
Site-level covariates:
point_id_year
ADAM01_A: 1
ADAM01_B: 1
ADAM02_A: 1
ADAM02_B: 1
ADAM03_A: 1
ADAM03_B: 1
(Other) :244
Yearly-site-level covariates:
year date time observer sky wind stream temp_c
A:300 Min. :16222 Min. :0.1868 Min. :1.000 5: 11 A:348 A:467 Min. : 1.50
B:450 1st Qu.:16248 1st Qu.:0.2285 1st Qu.:2.000 A:277 B:232 B:154 1st Qu.: 9.00
Median :16591 Median :0.2743 Median :3.000 B:175 C: 85 C: 80 Median :11.50
Mean :16458 Mean :0.2751 Mean :2.447 C:213 D: 50 D: 49 Mean :11.34
3rd Qu.:16604 3rd Qu.:0.3222 3rd Qu.:3.000 D: 44 E: 25 3rd Qu.:13.50
Max. :16613 Max. :0.4028 Max. :3.000 E: 30 F: 10 Max. :22.00
First I want to know: did I set the unmarked data frame up correctly given my objectives?
Next, I want to specify a global model, which I then want to run through the MuMIn() package’s dredge() function to derive the minimally adequate model involving only significant detection covariates as identified in this all-subset model selection process. Does anybody know if the dredge() function works with this particular model?
FYI, to reiterate my modeling objective: Given my best model, I want to derive abundance estimates at each individual site, corrected for imperfect detection, having taken into account temporary emigration between repeat surveys within each year.
Here is my model that I would kindly ask help setting up so as to achieve my modeling objective:
global <- gmultmix( ~1, ~1, ~date + time + sky + wind + stream + temp_c, data=zUMF, mixture = "NB")
I know that following the first tilde I should enter my abundance covariates (which I believe is analogous to the “state” side of a pcount() model). Given that I only want to correct for detection, I believe that I want to leave this null (~1)
I know that following the second tilde I should enter my availability covariates (which I believe are my interval counts, no?). Otherwise, what do I enter so as to model availability?
I know that following the third tilde I should enter my detection covariates.
Yet, the above model is all that I have been able to get to run. Otherwise, I haven’t had any success in correctly specifying the model. Given my objective, how should I specify this model?
Any and all help is much appreciated! Thanks!
-Tim
Dan Linden
Jan 22, 2016, 7:23:46 AM1/22/16
to unmarked
That looks correct, and it seems you've fit the model you want. I guess I'm not clear on what your question is specifically. Have you added covariates to other model processes and run into errors? What did those errors look like? Pretty sure dredge works with gmultmix but giving it a try would certainly clarify.
The best way to learn about these models is to try fitting different versions yourself, especially with simulated data. Richard has a ton of examples across the suite of unmarked models and most simulation techniques are applicable across multiple model types (they are more specific to R than unmarked). The help file for unmarkedFrameGMM has an illustration of covariates at each model process.
I would also suggest you grab Marc and Andy's new book, "Applied Hierarchical Modeling in Ecology". They cover everything you'd need to know about these models and they specifically discuss gmultmix in Section 7.5.3.
The best way to learn about these models is to try fitting different versions yourself, especially with simulated data. Richard has a ton of examples across the suite of unmarked models and most simulation techniques are applicable across multiple model types (they are more specific to R than unmarked). The help file for unmarkedFrameGMM has an illustration of covariates at each model process.
I would also suggest you grab Marc and Andy's new book, "Applied Hierarchical Modeling in Ecology". They cover everything you'd need to know about these models and they specifically discuss gmultmix in Section 7.5.3.
John Clare
Jan 26, 2016, 9:45:32 PM1/26/16
to unmarked
Just to follow up on what Dan said: yep, you have correctly identified tildes as abundance (null), availability (null), and detection (covariates). The actual counts should not be used as a covariate for availability, as this information should be completely contained within the removal detection history for each secondary period. You could use other covariates to model availability if thought plausible. One thing I would reiterate is that with the values of date being so large, the associated parameter estimate must be tiny relative to some of the others, and there may be some trouble finding an optima.
Tim Duclos
Jan 28, 2016, 2:03:42 PM1/28/16
to unmarked
John and Dan, I just want to thank you guys so much for taking the time to provide me with some great feedback. I'm still working on this, which goes without saying in the modeling and analysis world; I am confident that I will figure it out.
Dan, I wanna say that I did follow your suggestion and ordered Kery and Royle's new book "Applied Heirarchical Modeling in Ecology, V1". To say that this book is an incredible resource for a guy like me, is to make a huge understatement. It is written so very well and seemingly explains every detail of these models in terms that are extremely understandable (with step-by-step tutorials!)- a testament to the knowledge and expertise possessed by the authors as well as all the contributors associated with this guide. For me, this book could not have come at a more perfect time. Thanks for the heads up Dan! And thanks to all of you who contributed to the formation of these models!
Best,
Tim
Chris Merkord
Jan 28, 2016, 5:50:32 PM1/28/16
to unma...@googlegroups.com
I'll second Tim's comments about the new book. It makes the material very accessible.
I got the ebook, which on the one makes me sad because I like to sit back and admire the nice books on my shelf and feel the pages in my hands. Plus I sometimes remember things better when I can picture the pages they were on. But on the other hand, being able to search for keywords in the ebook is absolutely invaluable. Took me no time at all to find every page that had "c-hat" on it :-) (there are many).
If you don't have a copy yet, do yourself a favor and get one.
--
Christopher L. Merkord, Ph.D.
Postdoctoral Fellow, GSCE
South Dakota State University--
You received this message because you are subscribed to the Google Groups "unmarked" group.
To unsubscribe from this group and stop receiving emails from it, send an email to unmarked+u...@googlegroups.com.
For more options, visit https://groups.google.com/d/optout.
Reply all
Reply to author
Forward
0 new messages