Help needed in importing Brain tumor data into TVB GUI
29 views
Skip to first unread message
Maria Nazir
Dec 9, 2022, 1:14:22 PM12/9/22
to TVB Users
Dear all TVB community,
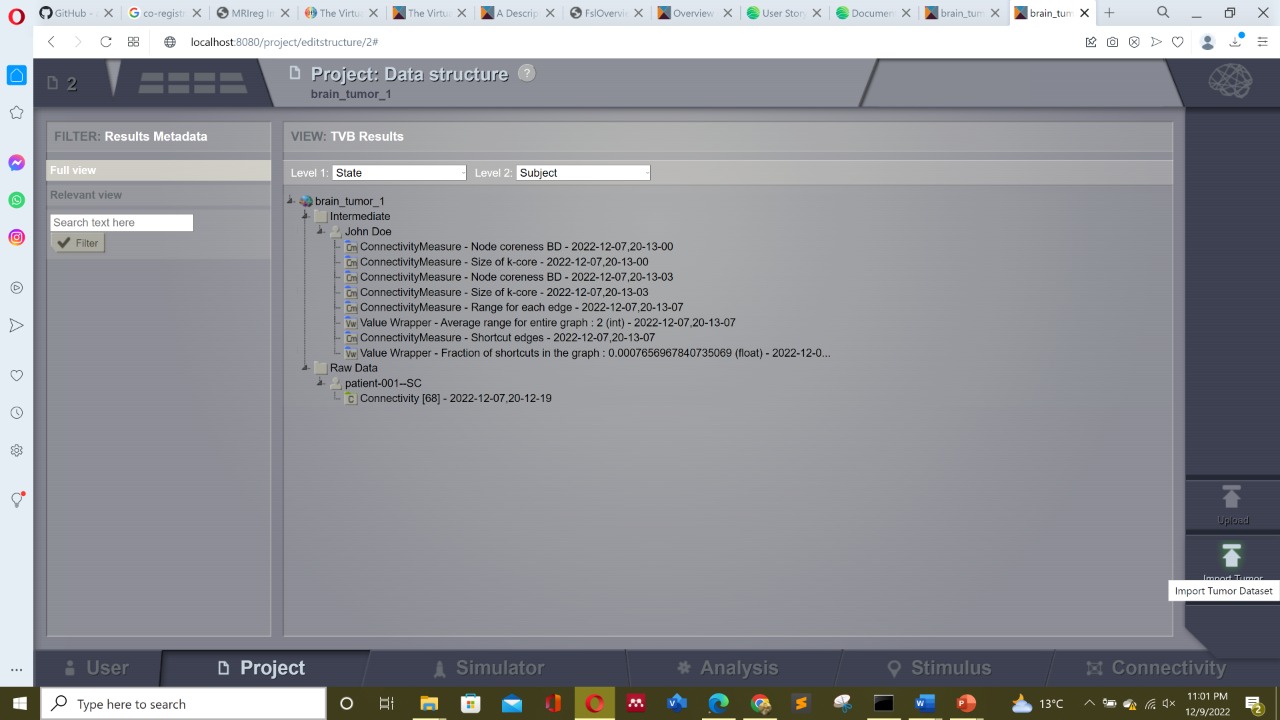
i am a new TVB user eagerly trying to work with it. My main work interest is in brain tumors however i am encountering problems while loading and uploading data into TVB GUI. My first question is that one can clearly see the icon of uploading tumor data as can be seen in the screenshot below on the right most bottom. But whenever i tried to load it , it never loads itself automatically and keep saying " it will take time to upload tumor data from ebrains etc " but never ever uploads it... can some one tell how can i do it ??

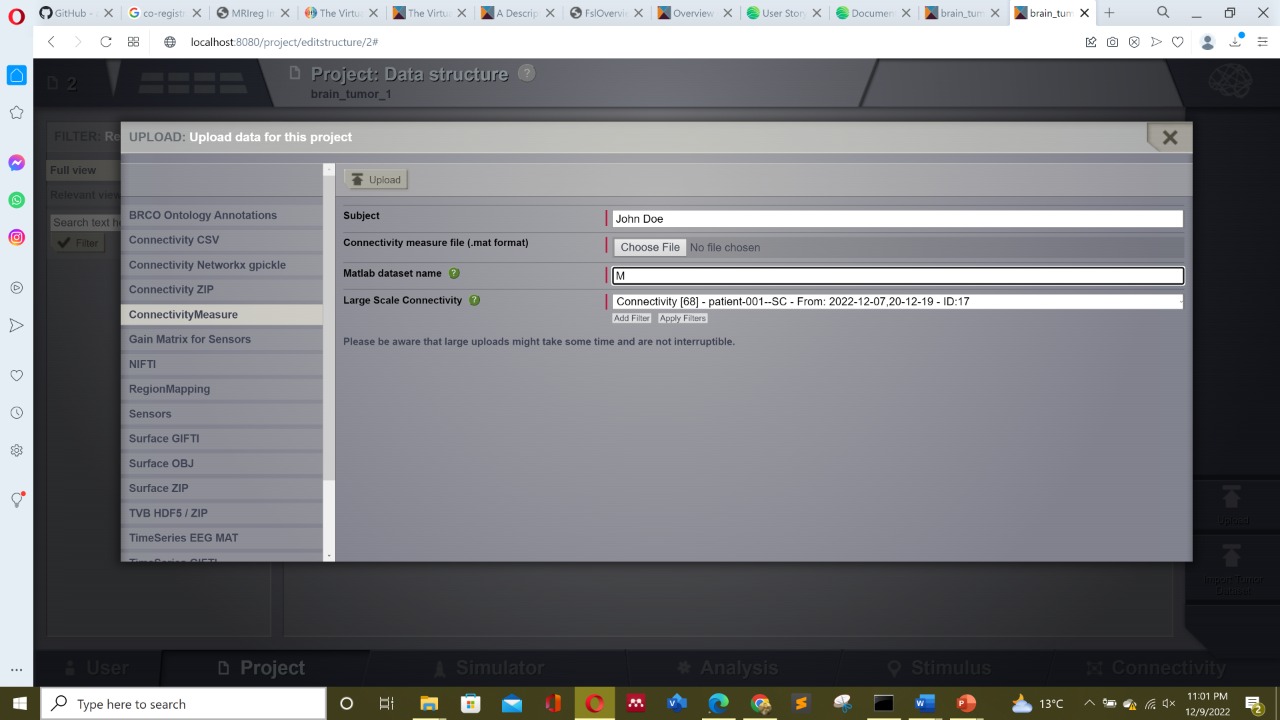
my second question is that whenever i try to load FC matrix from the pre-processed pre-operative brain tumor data , it never ever loads the matrix , may be becoz of the matlab directory. what should i enter in the place of matlab directory as can be seen in the figure below . Or is there another way to load FC matrix ?? please urgent help needed..

Lia Domide
Jan 2, 2023, 12:09:34 PM1/2/23
to TVB Users
Hi Marya,
EBRAINS made the Tumor Dataset private (it was available publicly until some time ago), thus our importer in TVB web interface is now broken.
We try to explain more the issue here: https://req.thevirtualbrain.org/browse/TVB-3057
We also have a code fix in this Pull Request: https://github.com/the-virtual-brain/tvb-root/pull/635 but it will become available only in the next releases of TVB (in a few weeks).
Until then, you can download the Tumor Dataset manually, and import each Structural Connectivity (SC.zip files) one by one (I suspect you already started that, from what I see in your screenshots).
Then, for the Functional Connectivities (FC.zip files), you can use the ConnectivityMeasure importer, as you started, but you should change the Matlab dataset name from the default 'M' (not valid in this case) into FC_cc_DK68
But, unfortunately that is not optimal, as it will result in 68 one dimensional vectors (one for each connectivity region), so you can only see the FC row by row.
Alternatively, you could try using TVB from Github (if you are comfortable to setup a Python env yourself), or I could provide you with some executable to run locally in your env and to made the folder compatible with your TVB version, if you say this is still actual for you; just give us a sign if we should proceed like this.
Best Regards,
Lia.
marya fahad
Jan 2, 2023, 11:38:38 PM1/2/23
to tvb-...@googlegroups.com
Thankyou so much Lia for the response. IF you could share with me the executables then it would give me a headstart and ill be highly obliged. Thankyou so much.
Waiting for a quick reply.
Sent from Mail for Windows
--
You received this message because you are subscribed to a topic in the Google Groups "TVB Users" group.
To unsubscribe from this topic, visit https://groups.google.com/d/topic/tvb-users/-fKhDBAJ6lU/unsubscribe.
To unsubscribe from this group and all its topics, send an email to tvb-users+...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/tvb-users/44b6b646-2e24-479e-94fb-d6d0a5da1c7dn%40googlegroups.com.
Lia Domide
Jan 9, 2023, 3:41:00 PM1/9/23
to TVB Users
Dear all,
In this version, the Import Tumor Dataset button to get the data from KB has been replaced with a regular uploader, on Project -> Upload -> Tumor Dataset
Before launching the new importer, you need to request and be granted access to EBRAINS KG dataset: https://search.kg.ebrains.eu/?category=Dataset&q=tumor#a696ccc7-e742-4301-8b43-d6814f3e5a44
In the time-windows they grant you access, you need to download the files locally, respecting the folders setup (it is close to BIDS, thus folders are important !!).
Then, you ZIP the top folder (one level above folders "sub-xxx"), and you pass it to TVB (ideally a local installation of tvb distribution, so that you do not infringe GDPR, or a secure installation, where you use TVB projects shared only with trusted other users).
For efficiency of the upload, you can also download + zip a subset of subjects at a time, from the full dataset in EBRAINS KG.
Have a good day,
TVB technical team.
Reply all
Reply to author
Forward
0 new messages
