Best multispectral to RGB conversion
jip
Some one asked me how I choose the matrix coef to convert a five multispectral image to a three (RGB) multispectral image. I feel that there is a best way to do that by maximising some distance in a color space.
How would you do that?
Examples of multispectral images are available here
Best Regard
Jean-Patrick Pommier
Zachary Pincus
> How would you do that?
> Examples of multispectral images are available here
More details on the input images would be helpful: are they the result of performing spectral unmixing, so each image represents the abundance of a particular fluorophore? Or are the images the raw results? The best approach would probably differ between the two options.
If the images are unmixed, then you can calculate the "true" spectrum at each pixel using the images and the emission spectra from the fluorophores. Then using these spectra, you could calculate the effective color in a number of ways, probably the simplest being a weighted average of the "red", "green", and "blue" regions of the spectra. You'd want to look at the psychophysics literature to figure out how best to do this, but I'm sure there are a ton of references online for converting a spectrum into an RGB color.
If the images are mixed (that is, raw), it's even easier: figure out the "apparent color" of the emission filter for each image (see below for sample code), and then colorize each image by the apparent color, and screen the images together. (Code for screening a set of images together is below, too -- 'multi_screen()')
Or, returning to the unmixed case, you could forget using the "natural" colors, and instead choose a set of colors maximally dissimilar from one another in some color space as the color for each fluorophore. This might make visualization simpler. You'd choose the colors (not sure how best), and then colorize the images and screen them together.
Zach
def screen(src, dst, max=255):
return max - (((max - src)*(max - dst))/max)
def multi_screen(images, max=255):
dst = images[0]
for image in images[1:]:
dst = screen(image, dst, max)
return dst
def wavelength_to_rgb(l):
# http://www.physics.sfasu.edu/astro/color/spectra.html
# So-called "Bruton's Algorithm"
assert (350 <= l <= 780)
if l < 440:
R = (440-l)/(440-350.)
G = 0
B = 1
elif l < 490:
R = 0
G = (l-440)/(490-440.)
B = 1
elif l < 510:
R = 0
G = 1
B = (510-l)/(510-490.)
elif l < 580:
R = (l-510)/(580-510.)
G = 1
B = 0
elif l < 645:
R = 1
G = (645-l)/(645-580.)
B = 0
else:
R = 1
G = 0
B = 0
if l > 700:
intensity = 0.3 + 0.7 * (780-l)/(780-700.)
elif l < 420:
intensity = 0.3 + 0.7 * (l-350)/(420-350.)
else:
intensity = 1
return intensity*numpy.array([R,G,B], dtype=float)
jip
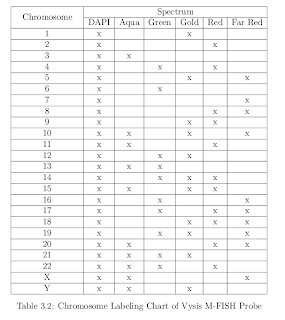
A given fluorochrome targets a subset of chromosomes, more informations can be found in:
Multiplex-Fluorescence In-Situ Hybridization
Chromosome Images
by
Hyo Hun Choi, B.S., M.S.
An example of integration times is [DAPI, Aqua,
Green, Gold, Red, Far Red] = [0.14, 6, 0.76, 6, 2.96, 1.4] seconds
He also gives an example of spectral mixing in the raw images:
A: S. Aqua
To perform "perfect" spectral unmixing, I suppose it's necessary to know the integration time, the characteristic of the microscope bandpass filters, the emission spectrum of each fluorochrome; but would it be possible to make some kind of "estimated" unmixing from only the images themselves?
Best regards
Jean-Pat
Zachary Pincus
> To perform "perfect" spectral unmixing, I suppose it's necessary to know the integration time, the characteristic of the microscope bandpass filters, the emission spectrum of each fluorochrome; but would it be possible to make some kind of "estimated" unmixing from only the images themselves?
Wait, are you trying to perform spectral unmixing (5 channels -> 25 channels, one per chromosome), or false coloring (5 channels -> 3 channel RGB)? The first email made it sound like the latter, but this one seems more on the former...
If you want to do unmixing, there's a large literature about the MFISH case specifically, and I think there are some pretty well-known methods at this point. Presumably it would help to know the integration times and spectral characteristics of the instrument / fluorophores, especially given the table you appended to the second email about the spectral mixing issues.
False-coloring is an easier subject. You'd just need to choose "target" RGB colors for each of the 5 raw images, colorize each image by that color, and screen them together. Most of the art is in choosing good "target" colors. You could choose based on the original colors ("aqua" etc.), or choose equidistant points on the hue-circle (with equivalent saturation, and brightness determined by the original grayscale pixel intensity).
Zach
On Mar 22, 2012, at 6:26 AM, jip wrote:
> Each image is a raw image of a given fluorochrome.
>
> A given fluorochrome targets a subset of chromosomes, more informations can be found in:
>
> Automatic Segmentation and Classification of
> Multiplex-Fluorescence In-Situ Hybridization
> Chromosome Images
> by
> Hyo Hun Choi, B.S., M.S.
> Choi gives an example of CCD integration times:
>
> An example of integration times is [DAPI, Aqua,
> Green, Gold, Red, Far Red] = [0.14, 6, 0.76, 6, 2.96, 1.4] seconds
>
> He also gives an example of spectral mixing in the raw images:
>
>
jip
Le jeudi 22 mars 2012 16:43:29 UTC+1, Zachary Pincus a écrit :
Jean-Pat,> To perform "perfect" spectral unmixing, I suppose it's necessary to know the integration time, the characteristic of the microscope bandpass filters, the emission spectrum of each fluorochrome; but would it be possible to make some kind of "estimated" unmixing from only the images themselves?
Wait, are you trying to perform spectral unmixing (5 channels -> 25 channels, one per chromosome), or false coloring (5 channels -> 3 channel RGB)? The first email made it sound like the latter, but this one seems more on the former...
Sorry, you're right, my first question was about the 5 ->3 channels issue.
If you want to do unmixing, there's a large literature about the MFISH case specifically, and I think there are some pretty well-known methods at this point. Presumably it would help to know the integration times and spectral characteristics of the instrument / fluorophores, especially given the table you appended to the second email about the spectral mixing issues.
False-coloring is an easier subject. You'd just need to choose "target" RGB colors for each of the 5 raw images, colorize each image by that color, and screen them together. Most of the art is in choosing good "target" colors. You could choose based on the original colors ("aqua" etc.),
I met that problem when I chose the matrix coef (with Stefan's code). The matrix was doing the job with:
'R': [0, 0, 0.05, 0.15, 0.8],
'V': [0.1, 0.8, 0.05, 0.05, 0],
'B': [0.8, 0.15, 0.05, 0, 0]}
or choose equidistant points on the hue-circle (with equivalent saturation, and brightness determined by the original grayscale pixel intensity).
*calculate saturation and brightness from gray scale s-aqua. (s1,b1)
*choose hue (a1)
Do the same for each fluo
Use the 5x3 hsb values to calculate a matrix to do the convertion 5 channels->3 RGB
Is it correct?
Jean-Pat
Zachary Pincus
> *calculate saturation and brightness from gray scale s-aqua. (s1,b1)
> *choose hue (a1)
> Do the same for each fluo
> Use the 5x3 hsb values to calculate a matrix to do the convertion 5 channels->3 RGB
>
> Is it correct?
Basically correct.
You could also just calculate brightness from the gray images, and leave the saturation constant.
Also, my suggestion was to use the "screen" operator to combine each image. The 5x3 matrix multiplication effectively makes 5 RGB images (one from each fluor), and then sums them together. In general, I find that "screen" works better than "sum" for making visually-appealing overlays of fluorescence channels.
For [0,1]-valued float images, the formula for screen-combining two images a and b is: a+b-ab. (http://en.wikipedia.org/wiki/Blend_modes#Screen )
Anyway, I'd try both sum and screen to see which looks best.
Zach