Interpreting "filter_stats" plot
27 views
Skip to first unread message
Victoria Glynn
Apr 14, 2022, 10:43:46 PM4/14/22
to poppr
Dear poppr community,
I hope this finds you well.
I wish to set my MLG cutoff threshold using filter_stats, but I am having issues wrapping my head around my current output.
I have a VCF file with SNP data that I converted into a SNPclone object, and then ran the following commands in R:
thresholds <- filter_stats(gl, distance = bitwise.dist, plot = TRUE)
thresholds
thresholds
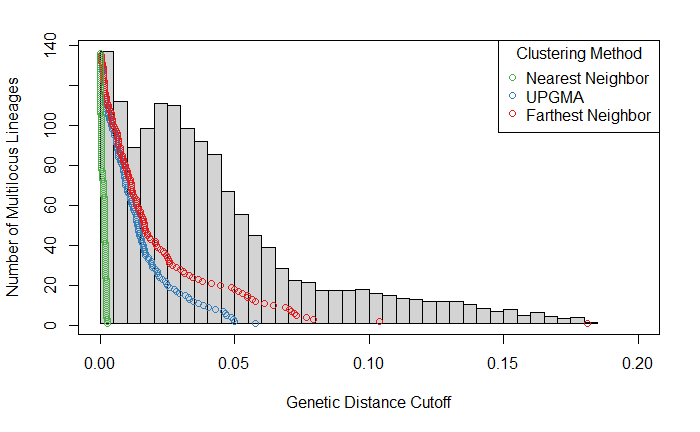
My plot looks as follow:

My questions are as follows:
- I am unsure if I am fully understanding how this plot is made and what it represents, as I would expect that as the genetic distance cut-off increases, the number of MLGs called decreases. However, there is an unexpected second peak in the histogram, close in size to the first one, at around a genetic distance of 0.025.
- From reading the online tutorial, I would interpret the grey histogram in the background to represent the total number of MLGs called at a given genetic distance cutoff. Is the histogram representing cumulative MLGs called across all three clustering methods, or a mean/median amount for all three clustering approaches?
Thank you in advance for your time and help. I really appreciate it.
Best regards,
Victoria
Zhian Kamvar
Apr 15, 2022, 12:36:03 PM4/15/22
to Victoria Glynn, poppr
Hello,
The histogram is an overlay visualisation of the distance matrix (e.g. hist(bitwise.dist(gl)) will give the same plot) and does not correspond to the y-axis (that's on me for not documenting this properly). An explanation of what it represents can be found in https://onlinelibrary.wiley.com/doi/10.1111/j.1365-294X.2007.03535.x#fu2.
It can be helpful in detecting the clonal peak (the first peak of the histogram) and determining a cutoff if there is no clear gap in the thresholds (shown as points on the graph).
The histogram is an overlay visualisation of the distance matrix (e.g. hist(bitwise.dist(gl)) will give the same plot) and does not correspond to the y-axis (that's on me for not documenting this properly). An explanation of what it represents can be found in https://onlinelibrary.wiley.com/doi/10.1111/j.1365-294X.2007.03535.x#fu2.
It can be helpful in detecting the clonal peak (the first peak of the histogram) and determining a cutoff if there is no clear gap in the thresholds (shown as points on the graph).
--
You received this message because you are subscribed to the Google Groups "poppr" group.
To unsubscribe from this group and stop receiving emails from it, send an email to poppr+un...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/poppr/a6c4f759-a68a-4965-8dce-86d0aad44335n%40googlegroups.com.
Victoria Glynn
Apr 28, 2022, 2:20:22 PM4/28/22
to poppr
Hello Zhian,
Thank you for your message and for providing further guidance on the plot. It is much appreciated.
Best regards,
Victoria
Reply all
Reply to author
Forward
0 new messages
