Plink2 PCA approx memory allocation
288 views
Skip to first unread message
Anthony Marcketta
May 21, 2020, 4:50:40 PM5/21/20
to plink2-users
Hi Cris,
What are the memory requirements for the PLINK2 PCA approx calculation?
My best guess currently is that Plink2 tries to allocate somewhere around:
Memory required = (Nsamples * Nvariants * NPCs * 4) bytes
Is this correct?
Thanks,
Tony
Christopher Chang
May 21, 2020, 6:15:50 PM5/21/20
to plink2-users
The leading terms are (Nvariants * 16 * NPCs^2) and (Nsamples * 16 * NPCs^2) bytes.
Anthony Marcketta
May 22, 2020, 8:56:24 AM5/22/20
to plink2-users
Oh I was way off. This is very helpful, thank you so much!
Out of curiosity, what are the memory requirements without the approx flag?
-Tony
Christopher Chang
May 22, 2020, 9:11:02 AM5/22/20
to plink2-users
Leading term is (Nsamples * Nsamples * 8) bytes.
Anthony Marcketta
Jul 16, 2020, 3:33:48 PM7/16/20
to plink2-users
Thanks!
I have another related question:
What is the approximate memory requirement for the plink2 --king-table function in terms of numbers of input samples, variants, and threads?
-Tony
Christopher Chang
Jul 17, 2020, 1:07:41 PM7/17/20
to plink2-users
That has a negligible memory requirement, compared to loading basic sample and variant info. --make-king-table automatically splits the overall computation into however many pieces are needed for each piece to fit in memory.
freeseek
Feb 21, 2023, 1:45:11 PM2/21/23
to plink2-users
Hi Christopher, I have been using plink2 --pca approx for a large biobank with the following parameters:
Nvariants = 287,959
Nsamples = 345,458
NPCs = 20
BED file size = 23.16GB
BIM file size = 0.01GB
FAM file size = 0.007GB
According to the suggested formula, the memory requirements would be:
(Nvariants + Nsamples) * 16 * NPCs^2 ~ 3.8GB
However, after running the job with ~7GB of memory and getting it repeatedly killed, I run it again monitoring what it is doing and, over the course of a three day run that successfully completed, the memory used by PLINK2 went from ~5.3GB to ~8.1GB.

This is more than double what estimated. Do you know what could be the reason and what would be the correct way to allocate memory for a VM that needs to run this type of computation?
Christopher Chang
Feb 21, 2023, 1:52:04 PM2/21/23
to plink2-users
Did you run with --memory?
If yes, and this was not actually limiting plink2's memory allocation, my best guess is that the linked linear algebra library is making large allocations of its own which I had not accounted for. I can investigate that if you provide enough details for me to reproduce the behavior.
freeseek
Feb 21, 2023, 2:05:55 PM2/21/23
to plink2-users
Yes, I had it run with --memory 6548 (VM had 7.2GB provided) and it died in ~20 minutes with the output below.

PLINK v2.00a3.5 AVX2 (9 Aug 2022) www.cog-genomics.org/plink/2.0/
(C) 2005-2022 Shaun Purcell, Christopher Chang GNU General Public License v3
Logging to FinnGenR11Affymetrix.log.
Options in effect:
--exclude range exclusion_regions.txt
--memory 6548
--out FinnGenR11Affymetrix
--pca 20 approx
--pgen FinnGenR11Affymetrix.prune.bed
--psam FinnGenR11Affymetrix.prune.fam
--pvar FinnGenR11Affymetrix.prune.bim
--threads 2
Start time: Tue Feb 14 22:19:49 2023
7450 MiB RAM detected; reserving 6548 MiB for main workspace.
Using up to 2 compute threads.
345458 samples (196050 females, 149408 males; 345458 founders) loaded from
FinnGenR11Affymetrix.prune.fam.
287959 variants loaded from FinnGenR11Affymetrix.prune.bim.
Note: No phenotype data present.
--exclude bed1: 3016 variants excluded.
Calculating allele frequencies... done.
284943 variants remaining after main filters.
Excluding 11246 variants on non-autosomes from PCA approximation.
Error: Out of memory. The --memory flag may be helpful.
(C) 2005-2022 Shaun Purcell, Christopher Chang GNU General Public License v3
Logging to FinnGenR11Affymetrix.log.
Options in effect:
--exclude range exclusion_regions.txt
--memory 6548
--out FinnGenR11Affymetrix
--pca 20 approx
--pgen FinnGenR11Affymetrix.prune.bed
--psam FinnGenR11Affymetrix.prune.fam
--pvar FinnGenR11Affymetrix.prune.bim
--threads 2
Start time: Tue Feb 14 22:19:49 2023
7450 MiB RAM detected; reserving 6548 MiB for main workspace.
Using up to 2 compute threads.
345458 samples (196050 females, 149408 males; 345458 founders) loaded from
FinnGenR11Affymetrix.prune.fam.
287959 variants loaded from FinnGenR11Affymetrix.prune.bim.
Note: No phenotype data present.
--exclude bed1: 3016 variants excluded.
Calculating allele frequencies... done.
284943 variants remaining after main filters.
Excluding 11246 variants on non-autosomes from PCA approximation.
Error: Out of memory. The --memory flag may be helpful.
Christopher Chang
Feb 21, 2023, 3:47:20 PM2/21/23
to plink2-users
Ok, this must be an overlooked part of the leading term, then. I will set up a job of this size tonight, look at all the memory allocations, and get back to you on what I missed.
Christopher Chang
Feb 21, 2023, 8:49:45 PM2/21/23
to plink2-users
Ok, revised major terms: (Nvariants * 16 * NPCs^2) + (Nsamples * 16 * NPCs^2) + (max(Nvariants, Nsamples) * 8 * NPCs^2) + (5760 * Nsamples) bytes. The third term was overlooked because it was based on a workspace size returned by the linear algebra library. The fourth term is driven by a high kPcaVariantBlockSize constant that I will consider halving in the next build.
Christopher Chang
Feb 21, 2023, 9:08:54 PM2/21/23
to plink2-users
another correction, the 8 in the third term should be a 16.
freeseek
Feb 22, 2023, 11:38:30 AM2/22/23
to plink2-users
This is great, thank you! Will this information be included in the PLINK2 documentation?
The successful run we had included the log below. In the "Projecting random vectors" line, 21 steps are described, rather than the number 20 of requested principal components. I assume this is part of how the algorithm works, but just to make sure, is NPCs in this case equal to 20 or to 21? Also, in the same successful run, I monitored the CPU usage after providing the VM with 2 CPUs and PLINK2 with 2 threads, and this is the CPU usage summary I have got:
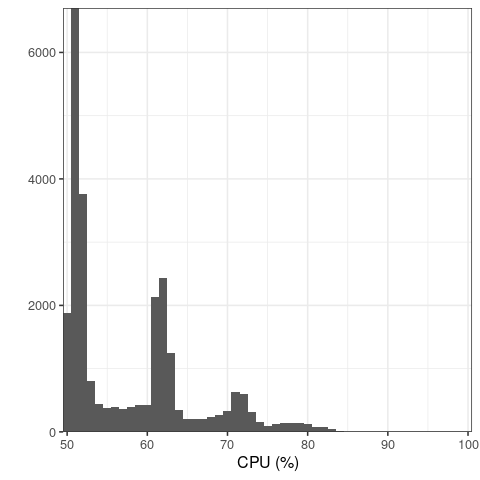
Most of the time PLINK2 is using 50% of the CPUs, that is 1 CPU in this example. So for the most part PLINK2 does not seem to be taking advantage of 2 threads. Is this a limitation of the algorithm? Would it make sense to provide even more threads for this type of computation?
PLINK v2.00a3.5 AVX2 (9 Aug 2022) www.cog-genomics.org/plink/2.0/
(C) 2005-2022 Shaun Purcell, Christopher Chang GNU General Public License v3
Logging to FinnGenR11Affymetrix.log.
Options in effect:
--exclude range exclusion_regions.txt
(C) 2005-2022 Shaun Purcell, Christopher Chang GNU General Public License v3
Logging to FinnGenR11Affymetrix.log.
Options in effect:
--exclude range exclusion_regions.txt
--memory 9946
--out FinnGenR11Affymetrix
--pca 20 approx
--pgen FinnGenR11Affymetrix.prune.bed
--psam FinnGenR11Affymetrix.prune.fam
--pvar FinnGenR11Affymetrix.prune.bim
--threads 2
Start time: Wed Feb 15 19:18:04 2023
11218 MiB RAM detected; reserving 9946 MiB for main workspace.
11218 MiB RAM detected; reserving 9946 MiB for main workspace.
Using up to 2 compute threads.
345458 samples (196050 females, 149408 males; 345458 founders) loaded from
FinnGenR11Affymetrix.prune.fam.
287959 variants loaded from FinnGenR11Affymetrix.prune.bim.
Note: No phenotype data present.
--exclude bed1: 3016 variants excluded.
Calculating allele frequencies... done.
284943 variants remaining after main filters.
Excluding 11246 variants on non-autosomes from PCA approximation.
Projecting random vectors (1 compute thread)... 21/21.
Computing SVD of Krylov matrix... done.
Recovering top PCs from range approximation... done.
--pca approx: Eigenvectors written to FinnGenR11Affymetrix.eigenvec , and
eigenvalues written to FinnGenR11Affymetrix.eigenval .
Computing SVD of Krylov matrix... done.
Recovering top PCs from range approximation... done.
--pca approx: Eigenvectors written to FinnGenR11Affymetrix.eigenvec , and
eigenvalues written to FinnGenR11Affymetrix.eigenval .
Christopher Chang
Feb 22, 2023, 11:59:51 AM2/22/23
to plink2-users
1. Okay, I will add this to the main documentation.
2. NPCs is 20. With that said, replacing all instances of NPCs^2 with NPCs * (NPCs + 1) gives you a more accurate formula, I just left out the +1 since it was not part of a leading term.
3. PLINK2's current implementation will only use more threads if you provide enough additional memory for them to work with. The per-thread workspace size is currently about (5760 * Nsamples) + (Nsamples * NPCs * (NPCs + 1) * 16) bytes.
Reply all
Reply to author
Forward
0 new messages
