Peptide confidence calculation
109 views
Skip to first unread message
Dmitry Abashkin
Jan 10, 2019, 7:25:02 AM1/10/19
to PeptideShaker
Hi! It seems unclear for me how peptide confidence is calculated. The PSM confidence is 1-PEP, which arises from the search procedure against the decoy database, but since we have prooved that the peptide presence is proven, what is this peptide confidence?
Thanks a lot!
Marc Vaudel
Jan 16, 2019, 7:29:14 AM1/16/19
to PeptideShaker
Hi Dmitry,
The confidence in the peptide identification is an aggregation of the confidence estimation of the different PSMs linking to the same peptide. The rationale is that multiple PSMs agreeing give you more confidence than a single PSM, this has mainly an influence on the ids with intermediate scores. Having the data summarized at the peptide level also allows estimating peptide-level FDR and better protein group scores. Finally, the presence of a peptide is never proven, that is why we try to give an estimate of the aggregated confidence.
Hope it helps, please don't hesitate if anything is unclear,
Marc
Le jeu. 10 janv. 2019 à 13:25, Dmitry Abashkin <dimab...@gmail.com> a écrit :
Hi! It seems unclear for me how peptide confidence is calculated. The PSM confidence is 1-PEP, which arises from the search procedure against the decoy database, but since we have prooved that the peptide presence is proven, what is this peptide confidence?Thanks a lot!
--
You received this message because you are subscribed to the Google Groups "PeptideShaker" group.
To unsubscribe from this group and stop receiving emails from it, send an email to peptide-shake...@googlegroups.com.
For more options, visit https://groups.google.com/d/optout.
Дмитрий абашкин
Jan 16, 2019, 1:58:15 PM1/16/19
to peptide...@googlegroups.com
Thank you, Marc. Now it's much more clear. But could you have a look at the added screenshot. Peptide number 38 (selected) had 39 not validated spectra (265 macthes before first decoy, all settings are default, FDR 1%). But the PSMs themselves have confidence around 80%, though they are "not validated" and the peptide confidence is 96%, unvalidated as well. How should I interpret the data? (In the particular case the experiment was a bit failure, but still I want to understand)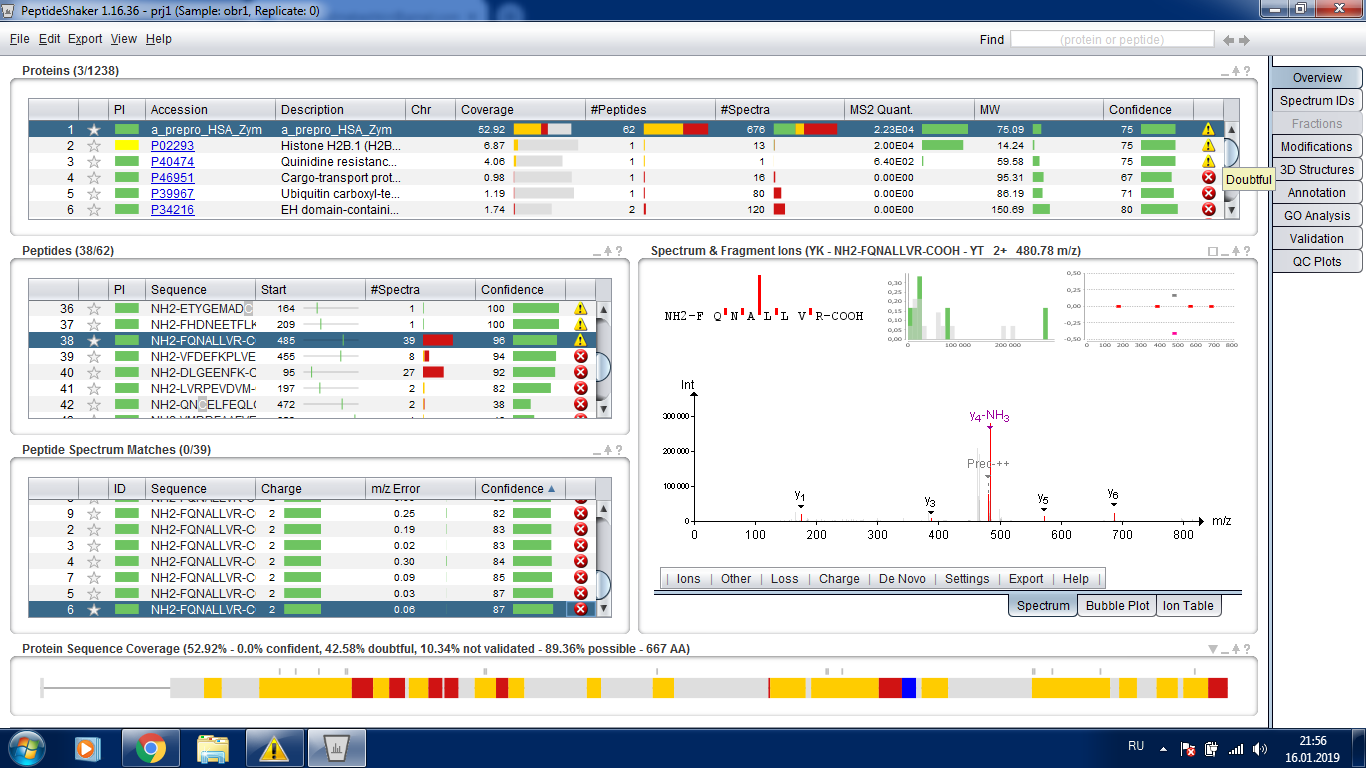

Marc Vaudel
Jan 18, 2019, 6:32:05 AM1/18/19
to PeptideShaker
Hi,
This is a very nice example: the PSMs have a confidence around 85%, and taking all PSMs with a confidence higher than this yields a dataset with FDR > 1%, this is why your PSMs are not validated. Taken all together, the 39 PSMs matching this peptide give you a confidence of 96% in the identification of the peptide. Taking all peptides with a confidence higher than 96% gives you an FDR < 1%, so the peptide is validated. It seems that you are right at the border, peptide 39 for this protein has a confidence of 94 and is not validated. As you can anticipate, with few peptides/proteins, the confidence estimation is not very reliable, so we need to take these results with a pinch of salt, this is why the tool is giving all these warning signs.
From the annotation of your spectrum, it seems that your MS2 accuracy is
very good, maybe being more stringent in MS2 tolerance would reduce the
prevalence of false positives?
Hope this clarifies what is happening under the hood, please do not hesitate if you find another counter-intuitive example like this one,
Marc
Дмитрий абашкин
Jan 18, 2019, 6:45:59 AM1/18/19
to peptide...@googlegroups.com
Thank you very much for superhelpful answer. Now I feel myself more confident with analyzing data ;) Actually the highlighted protein a_prepro_HSA_Zym was a bait in CoIP experiment, so I am perfectly sure that it was in the sample. But, unfortunately no other proteins-partners were found (this is the reply to your MS2 accuracy suggestion). As soon as we repeat that in different conditions, I can generate more questions. Thanks a lot, again
Marc Vaudel
Jan 18, 2019, 2:46:06 PM1/18/19
to PeptideShaker
Dear Dmitry,
Thanks for the kind words. CoIP are tricky, I often see the bait alone or half of Uniprot, which is not much more informative... Good luck with your experiments and please let us know if we can be of further help. Also, if you want to confirm the analysis with another software, I recommend giving a try to MaxQuant or Proline. Both tools will provide you with better quantitative estimates that will be very useful if you find other proteins :)
Best regards,
Marc
Reply all
Reply to author
Forward
0 new messages
