multi-variate normal sampling issue
14 views
Skip to first unread message
Dongchen Zhang
May 31, 2023, 3:23:36 PM5/31/23
to nimble-users
Hi guys,
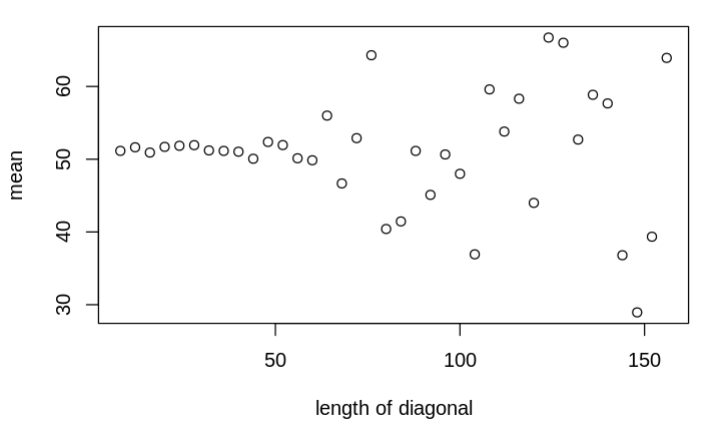
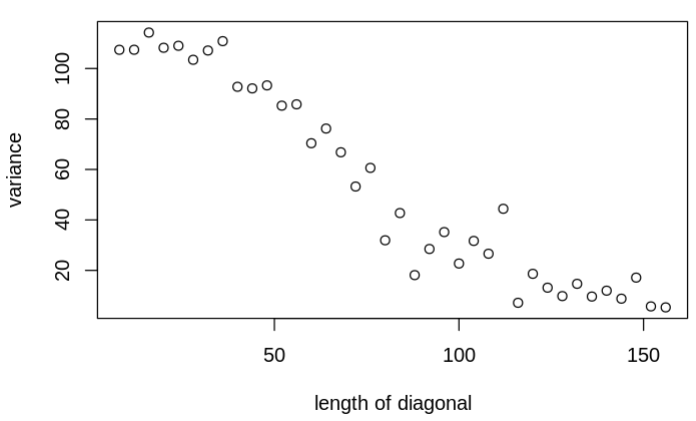
I am trying to use the mvnorm sampling to first sample x.mod from the given space (mean=mu.f; cov=Pf) and then create a likelihood comparing with the observations (y.censored) for those nodes that are not observed (with no observation) we used an H vector to only calculate the likelihood for those nodes that are observed. Currently, the Pf is just a diagonal matrix, each representing the variance of each node. However, with the increase of the Pf diagonal length (from 4 to 156), the unobserved sampled variable within the x.mod becomes unrealistic constrained (the variance drops from 110 to 10) and less converged, which should not be happening because there is no covariance amount those nodes. Interestingly, when I replace the mvnorm distribution with the sequential norm distribution using a for loop (see commented lines below, L3), the issue will be solved, making me wonder if there is something I should look into for the multi-variate normal sampling process. The model that I am using is as follows:
nimble.model <- nimbleCode({
# X model
# for (i in 1:N) {
# X.mod[i] ~ dnorm(mean = muf[i], sd = sqrt(pf[i, i]))
# }
X.mod[1:N] ~ dmnorm(mean = muf[1:N], cov = pf[1:N, 1:N])
for (i in 1:nH) {
tmpX[i] <- X.mod[H[i]]
Xs[i] <- tmpX[i]
}
## add process error to x model but just for the state variables that we have data and H knows who
X[1:YN] <- Xs[1:YN]
## Likelihood
y.censored[1:YN] ~ dmnorm(X[1:YN], prec = r[1:YN, 1:YN])
})
nimble.model <- nimbleCode({
# X model
# for (i in 1:N) {
# X.mod[i] ~ dnorm(mean = muf[i], sd = sqrt(pf[i, i]))
# }
X.mod[1:N] ~ dmnorm(mean = muf[1:N], cov = pf[1:N, 1:N])
for (i in 1:nH) {
tmpX[i] <- X.mod[H[i]]
Xs[i] <- tmpX[i]
}
## add process error to x model but just for the state variables that we have data and H knows who
X[1:YN] <- Xs[1:YN]
## Likelihood
y.censored[1:YN] ~ dmnorm(X[1:YN], prec = r[1:YN, 1:YN])
})
The figures that show the transitions when increasing the diagonal length are as follows:
I really appreciate your help!
Best,
Dongchen
Perry de Valpine
May 31, 2023, 5:07:27 PM5/31/23
to Dongchen Zhang, nimble-users
Dear Dongchen,
Thanks for the question. First, I confess that I am having a little bit of confusion following what you wrote, so I hope I understand your question. I think you are saying that the posterior variances of scalar elements of X.mod are smaller than they should be when you increase N.
The vector X.mod[1:N] will (by default) be updated by a Metropolis-Hastings sampler using multivariate normal proposals. For long vectors, this might not mix very well. The sampler is adaptive, meaning that the scale and correlation matrix of proposals goes through a self-tuning process. You can see parameters to control that in help(samplers) under the "RW_block" section. If the covariance pf is known, one idea would be to use that as the propCov (proposal covariance) and request adaptation only for the scale (a single factor that multiplies the entire proposal covariance matrix), by setting adaptive=TRUE and adaptScaleOnly=TRUE.
Another approach would be to sample X.mod as scalar elements or in smaller blocks. You can do that by modifying the default MCMC configuration. The User Manual Section 7.2 at r-nimble.org introduces how to do this.
HTH.
Perry
--
You received this message because you are subscribed to the Google Groups "nimble-users" group.
To unsubscribe from this group and stop receiving emails from it, send an email to nimble-users...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/nimble-users/733f0622-2bec-42e0-a82a-937a366590fdn%40googlegroups.com.
Dongchen Zhang
May 31, 2023, 5:12:24 PM5/31/23
to Perry de Valpine, nimble-users
Hi Perry,
Thank you so much for the responses, yes the question you asked was exactly what I was looking for and the solutions look promising, I will try both and let you know the progress!
Best,
Dongchen
Dongchen Zhang
May 31, 2023, 9:31:15 PM5/31/23
to Perry de Valpine, nimble-users
Hi Perry,
The first approach works! Thank you so much for the help! I also noticed that the chain is not well-mixed (see figure below), although the statistical results are correct (variance and mean values are almost identical to the Pf and mu.f). Do you have any idea about it?
Best,
Dongchen
Reply all
Reply to author
Forward
0 new messages
