Poisson lognormal convergence
Isaque Pim
Dear Nimble users,
I've been using Nimble to model case counts of the West African Ebola outbreak. I'm doing it in the same fashion people model the famous Scotland Lip Cancer dataset. Currently I'm having some trouble trying to make the model converge.
Actually, the basic Poisson lognormal model apparently presents no MCMC problems. When I add an ORLE term, autocorrelation spikes and the model shows poor efficiency (some betas presenting 0.1 ESS/s).
Here’s a sample of the attached code:
#Simple model, no ORLE
modelo1_simples <- nimbleCode({
for (i in 1:N){
y[i] ~ dpois(mu[i])
log(mu[i]) <- log(e[i]) + b0 + inprod(X[i,1:p],beta[1:p])
}
b0 ~ dnorm(0,0.001)
beta[1:p] ~ dmnorm(mean = m[1:p], cov = cov[1:p, 1:p])
})
# Model with ORLE
modelo2_ORLE <- nimbleCode({
for (i in 1:N){
y[i] ~ dpois(mu[i])
log(mu[i]) <- log(e[i]) + b0 + inprod(X[i,1:p],beta[1:p]) + u[i]
u[i] ~ dnorm(0, tau.b)
}
tau.b ~ dgamma(1,0.01)
b0 ~ dnorm(0,0.001)
beta[1:p] ~ dmnorm(mean = m[1:p], cov = cov[1:p, 1:p])
})
And I’m running quite long chains: 300,000 iter; 200,000 burnin; thinning = 10.
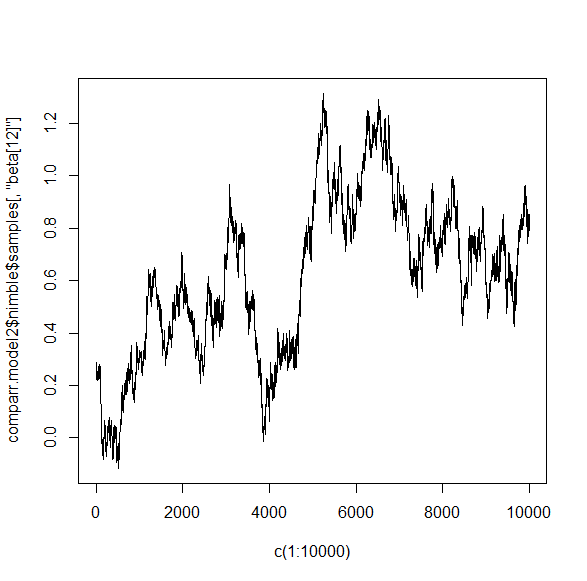
As another example, take a look at the traceplot for the same beta for each model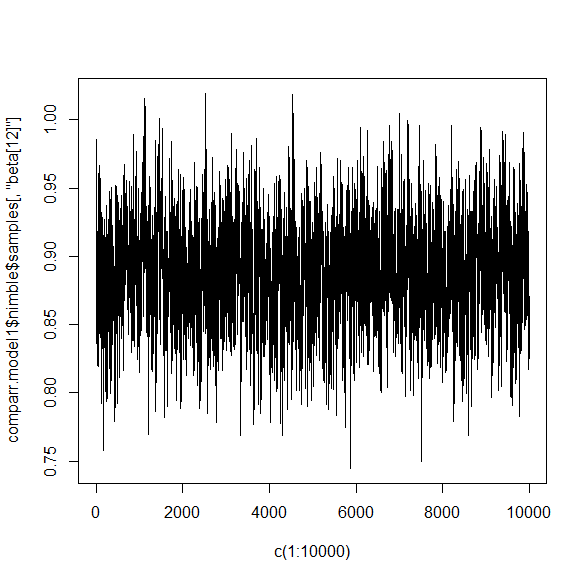
Chris Paciorek
--
You received this message because you are subscribed to the Google Groups "nimble-users" group.
To unsubscribe from this group and stop receiving emails from it, send an email to nimble-users...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/nimble-users/ad6d5409-4d54-4017-ae14-d617c607dfe9n%40googlegroups.com.
