BIOM <-> MetaPhlAn
911 views
Skip to first unread message
Sami Pietilä
Sep 28, 2013, 11:40:49 AM9/28/13
to metaphl...@googlegroups.com
Hi,
MetaPhlAn package is updated with biom2metaphlan.py, which can be used to convert OTU table from Qiime to MetaPhlAn relative abundance profile format, so that visualisation tools from MetaPhlAn pipeline can be used (hclust heatmap, lefse and krona).
The script is intended for early users who like to try new functionality (and report problems)
BR,
Sami
Zhuofei Xu
Jan 7, 2014, 8:00:13 AM1/7/14
to metaphl...@googlegroups.com
Hi All,
I met a problem when I converted a biom file into metaphlan profile format. Can anyone know which step is wrong? Thanks in advance!
The error information is below
python /home/zfxu/install/metaphlan/metaphlan1.7.3/conversion_scripts/biom2metaphlan.py -i otu_table_L6.biom -o otu_table_metaphlan.txt
Traceback (most recent call last):
File "/home/zfxu/install/metaphlan/metaphlan1.7.3/conversion_scripts/biom2metaphlan.py", line 416, in <module>
root = readBIOM(pars['i'])
File "/home/zfxu/install/metaphlan/metaphlan1.7.3/conversion_scripts/biom2metaphlan.py", line 217, in readBIOM
taxonomy = obs[2]["taxonomy"]
TypeError: 'NoneType' object has no attribute '__getitem__'
Best,I met a problem when I converted a biom file into metaphlan profile format. Can anyone know which step is wrong? Thanks in advance!
The error information is below
python /home/zfxu/install/metaphlan/metaphlan1.7.3/conversion_scripts/biom2metaphlan.py -i otu_table_L6.biom -o otu_table_metaphlan.txt
Traceback (most recent call last):
File "/home/zfxu/install/metaphlan/metaphlan1.7.3/conversion_scripts/biom2metaphlan.py", line 416, in <module>
root = readBIOM(pars['i'])
File "/home/zfxu/install/metaphlan/metaphlan1.7.3/conversion_scripts/biom2metaphlan.py", line 217, in readBIOM
taxonomy = obs[2]["taxonomy"]
TypeError: 'NoneType' object has no attribute '__getitem__'
--
You received this message because you are subscribed to the Google Groups "MetaPhlAn-users" group.
To unsubscribe from this group and stop receiving emails from it, send an email to metaphlan-use...@googlegroups.com.
For more options, visit https://groups.google.com/groups/opt_out.
--
Zhuofei Xu
Postdoc
Section of Microbiology
Department of Biology
University of Copenhagen
E-mail: zhuof...@gmail.com
Sami Pietilä
Jan 7, 2014, 3:15:39 PM1/7/14
to metaphl...@googlegroups.com
Hi Zhuofei,
I took a look at otu_table_Lx.biom files and they are somewhat different from otu_table.biom -files. Unfortunately the conversion script is currently unable to handle *Lx.biom -files. However, support for *Lx.biom files sounds like a good candidate for a new feature for next version of the script.
BR,
Sami
I took a look at otu_table_Lx.biom files and they are somewhat different from otu_table.biom -files. Unfortunately the conversion script is currently unable to handle *Lx.biom -files. However, support for *Lx.biom files sounds like a good candidate for a new feature for next version of the script.
BR,
Sami
Message has been deleted
Sx Chang
Nov 23, 2014, 10:46:07 AM11/23/14
to metaphl...@googlegroups.com
Dear Sami,
I am trying to convert a Biom file (MEGAN output) to metaphlan format.
Some errors happened as below:
root.maxdepth = max(root.maxdepth,len(taxonomy)-1)
TypeError: object of type 'NoneType' has no len()
If there are some format errors in this file? I hope you can provide a strict biom format example.
Thanks a lot.
With my best regards.
SX Chang
I am trying to convert a Biom file (MEGAN output) to metaphlan format.
Some errors happened as below:
Traceback (most recent call last):
File "biom2metaphlan.py", line 416, in <module>
root = readBIOM(pars['i'])
File "biom2metaphlan.py", line 218, in readBIOM
root.maxdepth = max(root.maxdepth,len(taxonomy)-1)
TypeError: object of type 'NoneType' has no len()
If there are some format errors in this file? I hope you can provide a strict biom format example.
Thanks a lot.
With my best regards.
SX Chang
Sami Pietilä
Nov 23, 2014, 11:13:51 AM11/23/14
to metaphl...@googlegroups.com
Hi,
biom2metaplan.py is written for BIOM files produced by Qiime. Unfortunately, Megan BIOM file might be somewhat different and thus be out of capabilities of this script. From the error, it looks as if the script can not find a field named "taxonomy" from the Megan BIOM file.
An example of a BIOM file written by Qiime can be found from: http://fmsc.utu.fi/metagenomics/ERP005367/run/otus/otu_table.biom.
Running a command "python biom2metaphlan.py -i otu_table.biom -o otu_table.metaplan" should convert the biom table to metaplan format when (biom-format 1.x library is required)
BR,
Sami
biom2metaplan.py is written for BIOM files produced by Qiime. Unfortunately, Megan BIOM file might be somewhat different and thus be out of capabilities of this script. From the error, it looks as if the script can not find a field named "taxonomy" from the Megan BIOM file.
An example of a BIOM file written by Qiime can be found from: http://fmsc.utu.fi/metagenomics/ERP005367/run/otus/otu_table.biom.
Running a command "python biom2metaphlan.py -i otu_table.biom -o otu_table.metaplan" should convert the biom table to metaplan format when (biom-format 1.x library is required)
BR,
Sami
Sx Chang
Nov 23, 2014, 9:18:45 PM11/23/14
to metaphl...@googlegroups.com
在 2014年11月24日星期一UTC+8上午12时13分51秒,Sami Pietilä写道:
Many Thanks.
Have a good day!
Have a good day!
touchp
Apr 28, 2015, 4:10:00 PM4/28/15
to metaphl...@googlegroups.com
Hi,
I got a problem when I use the script. (my biom file is a Qiime 1.9 output)
python biom2metaphlan.py -i otu_table.biom -o metaphlan.txt
Traceback (most recent call last):
File "biom2metaphlan.py", line 416, in <module>
root = readBIOM(pars['i'])
File "biom2metaphlan.py", line 211, in readBIOM
root.sampleIDs = list(table.SampleIds)
AttributeError: 'Table' object has no attribute 'SampleIds'
is it normal ?
I got a problem when I use the script. (my biom file is a Qiime 1.9 output)
python biom2metaphlan.py -i otu_table.biom -o metaphlan.txt
Traceback (most recent call last):
File "biom2metaphlan.py", line 416, in <module>
root = readBIOM(pars['i'])
root.sampleIDs = list(table.SampleIds)
AttributeError: 'Table' object has no attribute 'SampleIds'
is it normal ?
Sami Pietilä
Apr 29, 2015, 2:01:47 PM4/29/15
to metaphl...@googlegroups.com
Hi,
Qiime 1.9 has changed its BIOM format. biom2metaphlan.py does not understand the new format. I'll take a look if it is possible to update the script so that it would be able to read the new format.
It should also be possible to convert otu_table.biom into older format with biom -utils and then run biom2metaphlan.
BR,
Sami
Qiime 1.9 has changed its BIOM format. biom2metaphlan.py does not understand the new format. I'll take a look if it is possible to update the script so that it would be able to read the new format.
It should also be possible to convert otu_table.biom into older format with biom -utils and then run biom2metaphlan.
BR,
Sami
touchp
Apr 29, 2015, 4:24:13 PM4/29/15
to metaphl...@googlegroups.com
Hi,
Thanks for your reply but what is the command for biom -utils ?
Thanks for your reply but what is the command for biom -utils ?
George Weingart
Apr 29, 2015, 4:35:18 PM4/29/15
to touchp, metaphl...@googlegroups.com
Hi Patrick,
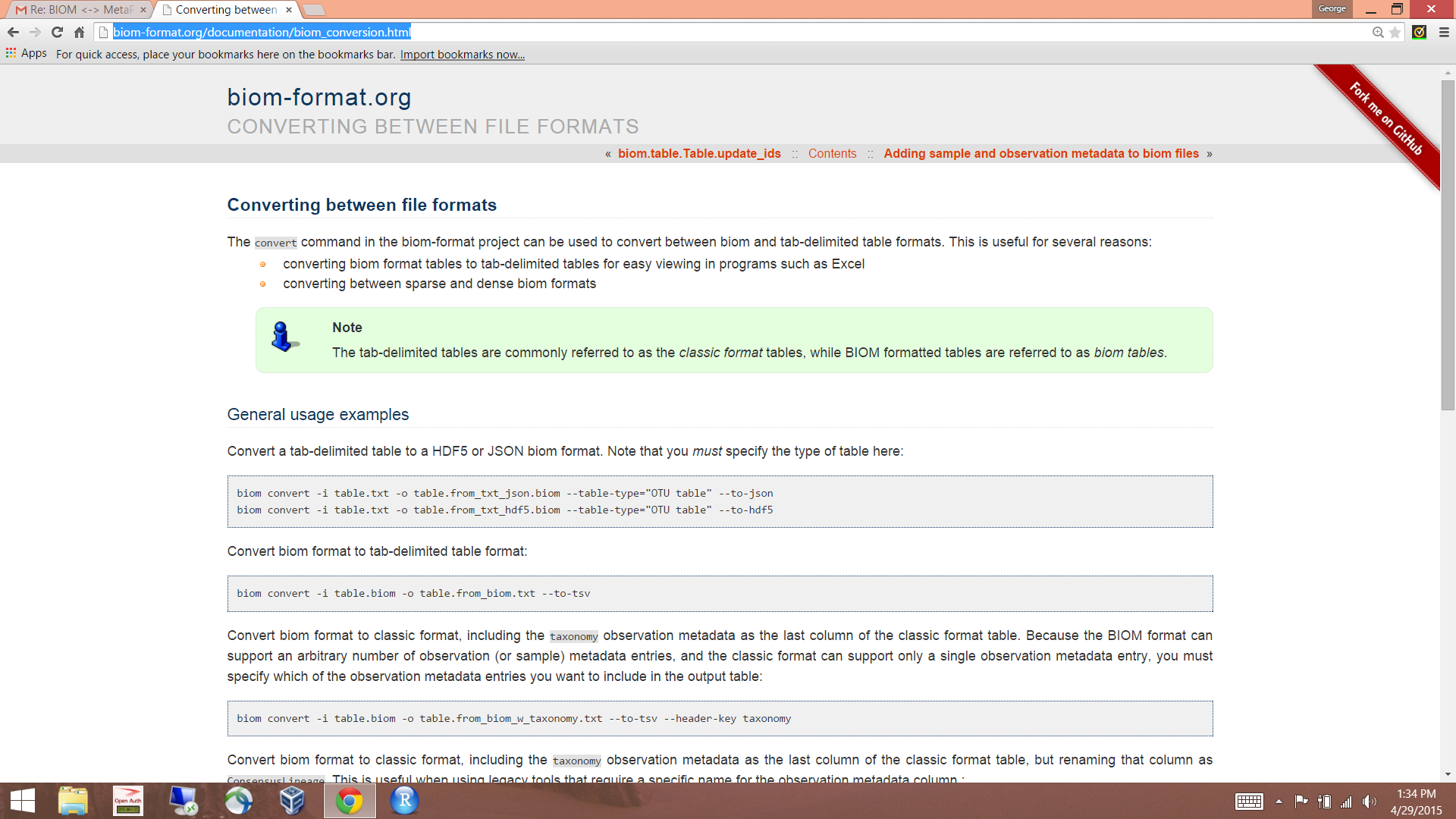
Best regards,
George Weingart, PhD
Huttenhower Lab

On Wed, Apr 29, 2015 at 1:24 PM, touchp <patrick...@gmail.com> wrote:
Hi,
Thanks for your reply but what is the command for biom -utils ?
--
You received this message because you are subscribed to the Google Groups "MetaPhlAn-users" group.
To unsubscribe from this group and stop receiving emails from it, send an email to metaphlan-use...@googlegroups.com.
For more options, visit https://groups.google.com/d/optout.
touchp
Apr 30, 2015, 12:10:40 PM4/30/15
to metaphl...@googlegroups.com
What is the version of QIIME that biom2metaphlan.py supports ?
Sami Pietilä
May 1, 2015, 7:12:02 AM5/1/15
to metaphl...@googlegroups.com
Older biom2metaphlan.py supports -> Qiime 1.8
If you want to test devel version of the script which should handle Qiime 1.9 format, it is available at:
https://ngs.utu.fi/files/biom/biom2metaphlan.py
If you want to test devel version of the script which should handle Qiime 1.9 format, it is available at:
https://ngs.utu.fi/files/biom/biom2metaphlan.py
touchp
May 1, 2015, 10:37:06 AM5/1/15
to metaphl...@googlegroups.com
Hi Sami. I got this error this time:
python =biom2metaphlan.py -i otu_table.biom -o otu_table_metahplan.txt
Traceback (most recent call last):
File "biom2metaphlan.py", line 461, in <module>
root, metaData = readBIOM(pars['i'])
File "biom2metaphlan.py", line 429, in readBIOM
samplemetadata = extractSampleMetadata(table)
File "biom2metaphlan.py", line 404, in extractSampleMetadata
keys = table.metadata(axis="sample")[0].keys()
python =biom2metaphlan.py -i otu_table.biom -o otu_table_metahplan.txt
Traceback (most recent call last):
root, metaData = readBIOM(pars['i'])
File "biom2metaphlan.py", line 429, in readBIOM
samplemetadata = extractSampleMetadata(table)
File "biom2metaphlan.py", line 404, in extractSampleMetadata
keys = table.metadata(axis="sample")[0].keys()
Sami Pietilä
May 2, 2015, 1:58:31 AM5/2/15
to metaphl...@googlegroups.com
Hi,
I forgot to mention that the new script version requires latest biom-format library (2.1). Can you check that you have the latest biom-format library?
Also the new script supports only biom files generated by Qiime 1.9.
BR,
Sami
I forgot to mention that the new script version requires latest biom-format library (2.1). Can you check that you have the latest biom-format library?
Also the new script supports only biom files generated by Qiime 1.9.
BR,
Sami
Reply all
Reply to author
Forward
0 new messages
