Discrepancy btwn voila view app and command line
52 views
Skip to first unread message
Yong shan
Jun 1, 2022, 4:03:17 AM6/1/22
to majiq_voila
Hi,
Thank you for creating this awesome software, it has been very useful to my lab recently.
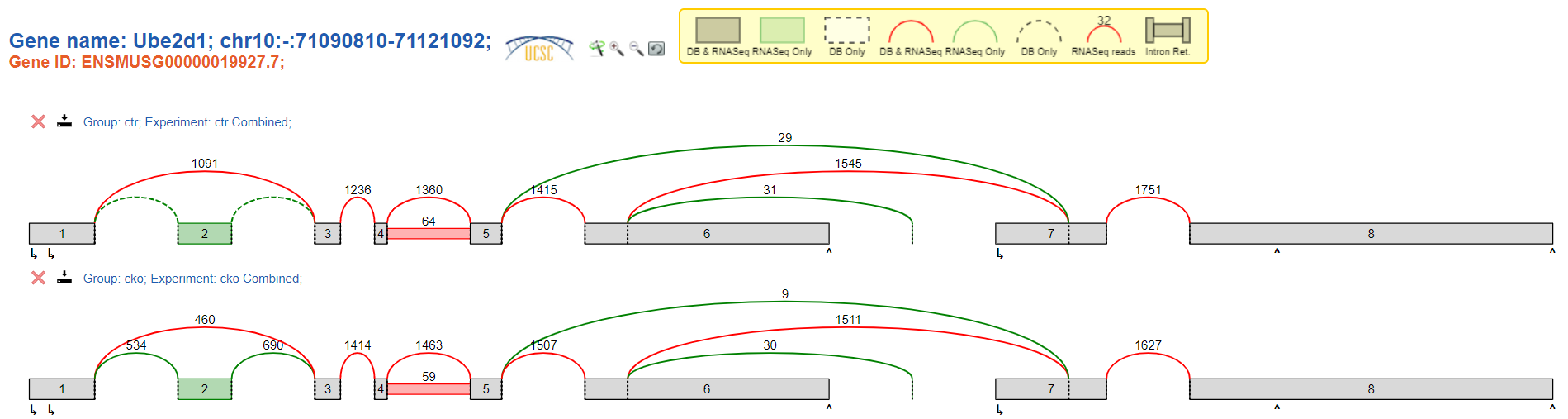
However, I realised that there is a discrepancy in read counts between voila viewing when using the GUI app (version: May 2019, I can't find the exact version number) and using the command line (version: 2.4.dev3+g85d07819), even when viewing the same gene for the exact same samples.
For example, for the same gene in the same set of samples,
Viewed using voila app:

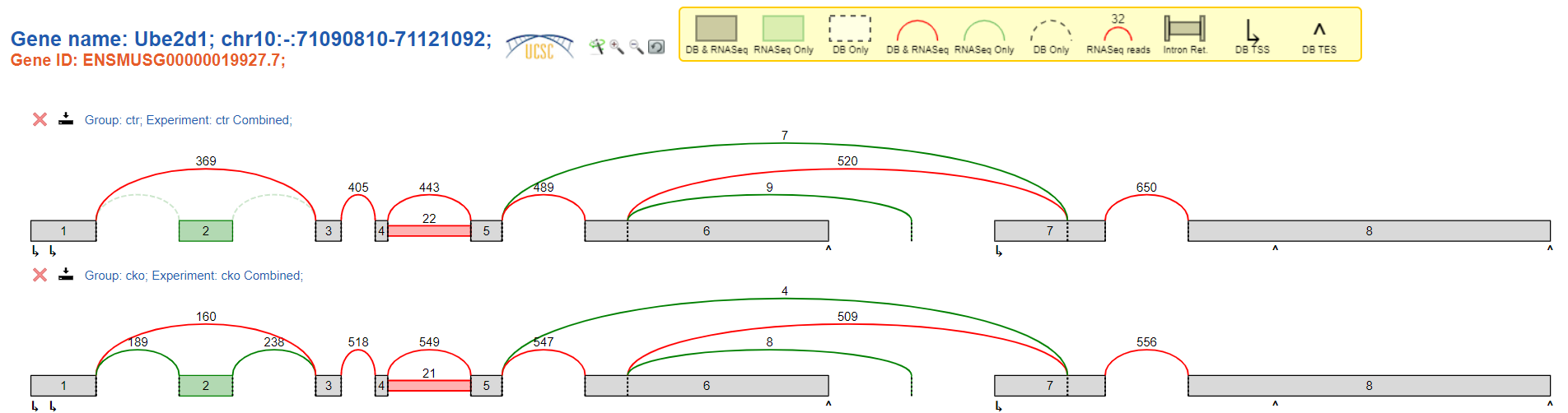
Viewed using terminal:

(command used: voila view ctr-cko.deltapsi.voila splicegraph.sql -p 5000 --host 0.0.0.0)
I have confirmed that I am looking at the same set of samples (3 controls + 3 treatment samples). It seems as though the terminal version is giving one-thirds as many reads as the app version.
Why is this the case? Am I using 2 different versions of voila? This discrepancy seems to be happening for all the genes in my samples.
I appreciate your time and help. Thank you very much!
Regards,
YS
Paul Jewell
Jun 2, 2022, 9:03:37 AM6/2/22
to majiq_voila
Hello YS,
Could you confirm which operating system you are using the app on?
There was a recent switch in the command line version which I believe is causing the change. Basically prior to 2.3, voila showed read counts 'combined' as the sum of all experiments in a group, and after, it showed the median red counts. The app version may be missing this update.
Yong shan
Jun 2, 2022, 9:56:09 PM6/2/22
to majiq_voila
Thank you for your quick reply! That explains it.
I am using Windows 10 64-bit operating system, x64-based processor
Reply all
Reply to author
Forward
0 new messages
