Mediation in Latent Growth Curve Models
843 views
Skip to first unread message
Judith B.
Dec 30, 2017, 2:43:34 AM12/30/17
to lavaan
I'm a complete newbie to R, and learning as I'm going. I haven't had much luck trying to find syntax to conduct a mediation analysis in latent growth curve modeling framework. Using lavaan I've been able to run mediation and latent growth curve models separately, but my attempts to to integrate those two has failed.
I would appreciate it if someone was willing to share some code they have. I have one predictor and one mediator variable (both measured at time 1), and a dependent variable measured at time 1, time 2, and time 3.
Thanks!
I would appreciate it if someone was willing to share some code they have. I have one predictor and one mediator variable (both measured at time 1), and a dependent variable measured at time 1, time 2, and time 3.
Thanks!
Terrence Jorgensen
Jan 5, 2018, 7:54:42 AM1/5/18
to lavaan
I've been able to run mediation and latent growth curve models separately, but my attempts to to integrate those two has failed.
My advice would be to draw a path diagram of your hypothesized model. If you can draw it, you can translate it into syntax using "~" for regression paths (or "=~" for factor loadings) and "~~" for (co)variances.
I would appreciate it if someone was willing to share some code they have. I have one predictor and one mediator variable (both measured at time 1), and a dependent variable measured at time 1, time 2, and time 3.
So your dependent variables in the mediation model are the growth factors (latent intercept and slope)? Simply regress those onto your mediator, and regress the mediator onto your predictor.
Terrence D. Jorgensen
Postdoctoral Researcher, Methods and Statistics
Research Institute for Child Development and Education, the University of Amsterdam
UvA web page: http://www.uva.nl/profile/t.d.jorgensen
Judith B.
Jan 9, 2018, 6:28:55 PM1/9/18
to lavaan
Hi Terrence,
thanks! I think I fixed my mistakes, including several error messages I received. My model looks like this.
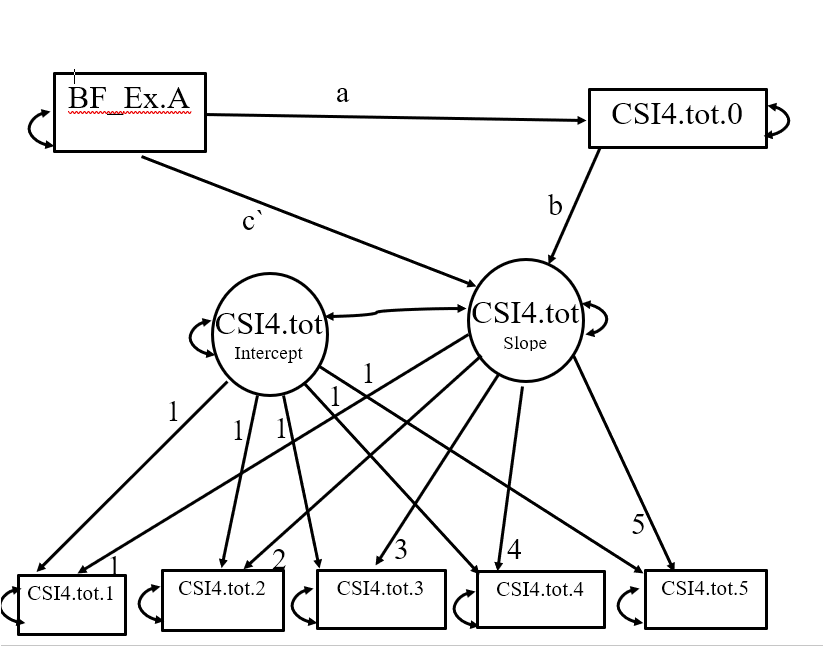
Essentially, I'm examining whether CSI4.tot at time 0 mediates the association between "BF_Ex.A" and subsequent changes in CSI4.tot at times 1 through 5.
Here is my syntax:
I can run this model without getting any error messages, but I am not absolutely confident that my code is accurate. Some feedback would be appreciated.
Thanks!
thanks! I think I fixed my mistakes, including several error messages I received. My model looks like this.
Essentially, I'm examining whether CSI4.tot at time 0 mediates the association between "BF_Ex.A" and subsequent changes in CSI4.tot at times 1 through 5.
Here is my syntax:
mediation.model <- '
#latent growth variables
#DV intercept
i =~ 1*CSI4.tot.1 + 1*CSI4.tot.2 + 1*CSI4.tot.3 + 1*CSI4.tot.4 + 1*CSI4.tot.5
#DV slope
s =~ 1*CSI4.tot.1 + 2*CSI4.tot.2 + 3*CSI4.tot.3 + 4*CSI4.tot.4 + 5*CSI4.tot.5
#Specifying intercept for CSI4.tot.0
CSI4.tot.0 ~ 1
#direct effect
i ~ c*BF_Ex.A
s ~ c*BF_Ex.A
# mediator
CSI4.tot.0 ~ a*BF_Ex.A
i ~ b*CSI4.tot.0
s ~ b*CSI4.tot.0
# indirect effect (a*b)
ab := a*b
# total effect
total := c + (a*b)
'
fit.growth<-growth(mediation.model, data = dset, se = "bootstrap")
summary(fit.growth)
I can run this model without getting any error messages, but I am not absolutely confident that my code is accurate. Some feedback would be appreciated.
Thanks!
Terrence Jorgensen
Jan 10, 2018, 2:44:43 PM1/10/18
to lavaan
You should also estimate the intercept of i and of s, but the growth() function should add those parameters automatically. You also need to use different labels for the effects on i and on s (using b and c for both constrains those effects to be equal, which is not what you want to do). You can use {ib, ic} for effects on i, and {sb, sc} for effects on s.
Judith B.
Jan 10, 2018, 3:34:41 PM1/10/18
to lavaan
Thanks again, Terrence.
Just to be sure, given that the growth() function automatically estimates the latent variable intercepts, there is nothing else I need to do, correct?
Does this code and output look accurate then?
Code:
Just to be sure, given that the growth() function automatically estimates the latent variable intercepts, there is nothing else I need to do, correct?
Does this code and output look accurate then?
Code:
mediation.model <- '
#latent growth variables (note: only DV is growth factor in this model)
#DV intercept
i =~ 1*CSI4.tot.1 + 1*CSI4.tot.2 + 1*CSI4.tot.3 + 1*CSI4.tot.4 + 1*CSI4.tot.5
#DV slope
s =~ 1*CSI4.tot.1 + 2*CSI4.tot.2 + 3*CSI4.tot.3 + 4*CSI4.tot.4 + 5*CSI4.tot.5
#Specifying intercept for CSI4.tot.0
CSI4.tot.0 ~ 1
#direct effect
i ~ {ic}*BF_Ex.A
s ~ {sc}*BF_Ex.A
# mediator
CSI4.tot.0 ~ a*BF_Ex.A
i ~ {ib}*CSI4.tot.0
s ~ {sb}*CSI4.tot.0
# indirect effect (a*{sb})
ab := a*{sb}
# total effect
total := {sc} + (a*{sb})
'
fit.growth<-growth(mediation.model, data = dset, se = "bootstrap")
summary(fit.growth)
Output:
lavaan (0.5-23.1097) converged normally after 47 iterations Used Total Number of observations 165 600 Estimator ML Minimum Function Test Statistic 55.211 Degrees of freedom 16 P-value (Chi-square) 0.000 Parameter Estimates: Information Observed Standard Errors Bootstrap Number of requested bootstrap draws 1000 Number of successful bootstrap draws 1000 Latent Variables: Estimate Std.Err z-value P(>|z|) i =~ CSI4.tot.1 1.000 CSI4.tot.2 1.000 CSI4.tot.3 1.000 CSI4.tot.4 1.000 CSI4.tot.5 1.000 s =~ CSI4.tot.1 1.000 CSI4.tot.2 2.000 CSI4.tot.3 3.000 CSI4.tot.4 4.000 CSI4.tot.5 5.000 Regressions: Estimate Std.Err z-value P(>|z|) i ~ BF_Ex.A (ic) 0.097 0.077 1.251 0.211 s ~ BF_Ex.A (sc) -0.005 0.022 -0.210 0.834 CSI4.tot.0 ~ BF_Ex.A (a) -0.034 0.089 -0.381 0.703 i ~ CSI4.tt.0 (ib) 0.887 0.054 16.485 0.000 s ~ CSI4.tt.0 (sb) -0.038 0.017 -2.213 0.027 Covariances: Estimate Std.Err z-value P(>|z|) .i ~~ .s -0.853 0.253 -3.365 0.001 Intercepts: Estimate Std.Err z-value P(>|z|) .CSI4.tot.0 9.028 0.657 13.737 0.000 .CSI4.tot.1 0.000 .CSI4.tot.2 0.000 .CSI4.tot.3 0.000 .CSI4.tot.4 0.000 .CSI4.tot.5 0.000 .i -0.327 0.741 -0.441 0.659 .s 1.122 0.230 4.878 0.000 Variances: Estimate Std.Err z-value P(>|z|) .CSI4.tot.1 2.569 0.555 4.631 0.000 .CSI4.tot.2 2.977 0.608 4.895 0.000 .CSI4.tot.3 4.290 0.811 5.293 0.000 .CSI4.tot.4 2.018 0.305 6.624 0.000 .CSI4.tot.5 0.728 0.489 1.487 0.137 .CSI4.tot.0 13.276 1.438 9.230 0.000 .i 7.605 1.284 5.924 0.000 .s 0.534 0.082 6.524 0.000 Defined Parameters: Estimate Std.Err z-value P(>|z|) ab 0.001 0.004 0.352 0.725 total -0.003 0.022 -0.150 0.881
Thanks!
Terrence Jorgensen
Jan 11, 2018, 12:31:09 PM1/11/18
to lavaan
Does this code and output look accurate then?
Those curly braces should not be there, and you could also calculate the indirect effect on i, but it looks like it worked (all the expected estimates are in the output).
ahmad
Feb 23, 2026, 9:03:32 AMFeb 23
to lavaan
Hi, following up on this point, I’m wondering how we can extend this model to a moderated mediation framework using multiple-group SEM. Specifically, if we want to examine the moderating effects of sex on the direct, indirect, and total effects in this latent growth mediation model, should we constrain any parameters in the latent growth model portion? Or can we treat this model as a standard mediation model and simply test the moderating role of sex across groups?
Thank,
A
Terrence Jorgensen
Mar 26, 2026, 5:12:16 AMMar 26
to lavaan
I’m wondering how we can extend this model to a moderated mediation framework using multiple-group SEM. Specifically, if we want to examine the moderating effects of sex on the direct, indirect, and total effects in this latent growth mediation model, should we constrain any parameters in the latent growth model portion?
Such as? If you are talking about the (co)variances of the growth factors (or the residual variances of the repeated measures), then I would say an advantage of the multigroup approach is that you do not have to assume homogeneity of variances across groups. Certainly, if it is interesting to test a hypothesis of equal variance, you can impose constraints to compare models with a LRT, but it sounds like you are just interested in comparing the regression slopes.
Terrence D. Jorgensen (he, him, his)
Assistant Professor, Methods and Statistics
Assistant Professor, Methods and Statistics
Research Institute for Child Development and Education, the University of Amsterdam
Reply all
Reply to author
Forward
0 new messages
