Substructure Highlighting Help
31 views
Skip to first unread message
Ryan Lougee
Jul 27, 2020, 11:28:29 AM7/27/20
to indigo-general
Hi All,
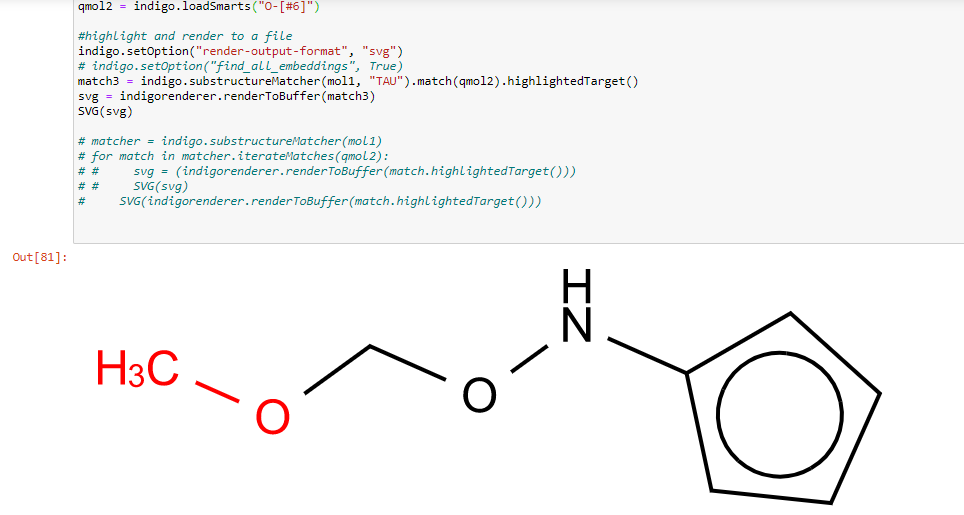
I was interested in making an open source tool for substructure highlighting. I'm not sure there is a great way to do this currently.
So far I am working in python, and I am able to highlight a single substructure from a Smarts:
However, I have not been able to achieve a couple goals:
1) highlight multiple instances on the same substructure
2) highlight different substructures within the same molecule
I'm not too familiar with C++, but it looks like I could just switch the "find_all_embeddings" bool to True in "molecule/src/molecule_tautomer_substructure_matcher.cpp" or a related directory. That might achieve my first goal but I'm not sure how I might accomplish the second.
So, is it possible to do these things in python or should I make a new branch and start learning C++?
Thanks,
-Ryan
Reply all
Reply to author
Forward
0 new messages
