Access certain group..
211 views
Skip to first unread message
Elsaid Younes
Aug 16, 2021, 5:36:14 AM8/16/21
to hoomd...@googlegroups.com
Dear all,
data1 = np.linspace([-451.0,-451.0,-451.0], [451.0,451.0,451.0], 2940)
np.random.shuffle(data1[:,0])
np.random.shuffle(data1[:,1])
I wonder How can I access particles with type'B'
GSD file:
data1 = np.linspace([-451.0,-451.0,-451.0], [451.0,451.0,451.0], 2940)
np.random.shuffle(data1[:,0])
np.random.shuffle(data1[:,1])
data2 = np.linspace([-376.0,-376.0,-376.0], [376.0,376.0,376.0], 21)
np.random.shuffle(data2[:,0])
np.random.shuffle(data2[:,1])
result = [*data1,*data2]
np.random.shuffle(data2[:,0])
np.random.shuffle(data2[:,1])
result = [*data1,*data2]
s = gsd.hoomd.Snapshot()
s.particles.N = 2961
s.particles.typeid = [0]*2940 + [1]*21
s.particles.types = ['A']*2940 + ['B']*21
s.particles.diameter = [4.0]*2940 + [150.0]*21
s.configuration.box = [906.0,906.0,906.0,0,0,0]
s.particles.charge = [+1]*2940 + [-140]*21
s.particles.position = result
with gsd.hoomd.open(name='file.gsd', mode='wb') as f:
f.append(s)
s.particles.N = 2961
s.particles.typeid = [0]*2940 + [1]*21
s.particles.types = ['A']*2940 + ['B']*21
s.particles.diameter = [4.0]*2940 + [150.0]*21
s.configuration.box = [906.0,906.0,906.0,0,0,0]
s.particles.charge = [+1]*2940 + [-140]*21
s.particles.position = result
with gsd.hoomd.open(name='file.gsd', mode='wb') as f:
f.append(s)
Best regards,
Elsaid Mohamed
Eric Irrgang
Aug 16, 2021, 8:16:42 AM8/16/21
to hoomd...@googlegroups.com
Hi Elsaid.
1. There is a logic error in the script you share. Per https://gsd.readthedocs.io/en/stable/hoomd-examples.html#define-a-snapshot, the `types` field is supposed to be a list of types, such that the indices of the list correspond to the integers used in the `typeid` field.
2. You would find particles of a given type by looking for particles with a given typeid. Python has several ways to do this for lists, but since GSD provides a NumPy interface, you could do something like
>>> s.particles.typeid = [0]*2940 + [1]*21
>>> s.particles.types = ['A', 'B']
>>> with gsd.hoomd.open(name='file.gsd', mode='rb') as t: frame = t.read_frame(0)
...
>>> frame.particles.position[:].shape
(2961, 3)
>>> frame.particles.typeid[:] == 0
array([ True, True, True, ..., False, False, False])
>>> frame.particles.position[frame.particles.typeid[:] == 0].shape
(2940, 3)
>>> for pos in frame.particles.position[frame.particles.typeid == frame.particles.types.index('B')]: # Do something...
Note that that last line only works if you fix the `types` as noted above.
See https://numpy.org/doc/stable/reference/arrays.indexing.html#advanced-indexing (or just https://www.w3schools.com/python/python_lists_comprehension.asp)
Best,
Eric
> --
> You received this message because you are subscribed to the Google Groups "hoomd-users" group.
> To unsubscribe from this group and stop receiving emails from it, send an email to hoomd-users...@googlegroups.com.
> To view this discussion on the web visit https://groups.google.com/d/msgid/hoomd-users/CAO9cOeB6SRTPbEE9FdKvFV-f_WGT4jXLmFOiPtAwTsJr%2BaUyUA%40mail.gmail.com.
1. There is a logic error in the script you share. Per https://gsd.readthedocs.io/en/stable/hoomd-examples.html#define-a-snapshot, the `types` field is supposed to be a list of types, such that the indices of the list correspond to the integers used in the `typeid` field.
2. You would find particles of a given type by looking for particles with a given typeid. Python has several ways to do this for lists, but since GSD provides a NumPy interface, you could do something like
>>> s.particles.typeid = [0]*2940 + [1]*21
>>> with gsd.hoomd.open(name='file.gsd', mode='wb') as f: f.append(s)
...
>>> with gsd.hoomd.open(name='file.gsd', mode='rb') as t: frame = t.read_frame(0)
...
>>> frame.particles.position[:].shape
(2961, 3)
>>> frame.particles.typeid[:] == 0
array([ True, True, True, ..., False, False, False])
>>> frame.particles.position[frame.particles.typeid[:] == 0].shape
(2940, 3)
>>> for pos in frame.particles.position[frame.particles.typeid == frame.particles.types.index('B')]: # Do something...
Note that that last line only works if you fix the `types` as noted above.
See https://numpy.org/doc/stable/reference/arrays.indexing.html#advanced-indexing (or just https://www.w3schools.com/python/python_lists_comprehension.asp)
Best,
Eric
> You received this message because you are subscribed to the Google Groups "hoomd-users" group.
> To unsubscribe from this group and stop receiving emails from it, send an email to hoomd-users...@googlegroups.com.
> To view this discussion on the web visit https://groups.google.com/d/msgid/hoomd-users/CAO9cOeB6SRTPbEE9FdKvFV-f_WGT4jXLmFOiPtAwTsJr%2BaUyUA%40mail.gmail.com.
Elsaid Younes
Aug 16, 2021, 4:00:02 PM8/16/21
to hoomd...@googlegroups.com
Dear all,
GSD file:
s = gsd.hoomd.Snapshot()
s.particles.N = 2961
s.particles.typeid = [0]*2940 + [1]*21
s.particles.N = 2961
s.particles.typeid = [0]*2940 + [1]*21
s.particles.types = ['A','B']
s.particles.diameter = [4.0]*2940 + [150.0]*21
s.configuration.box = [906.0,906.0,906.0,0,0,0]
s.particles.charge = [+1]*2940 + [-140]*21
s.particles.position = result
with gsd.hoomd.open(name='file.gsd', mode='wb') as f:
f.append(s)
Simulation script:
import hoomd
import freud
import gsd.hoomd
import numpy as np
import matplotlib.pyplot as plt
import hoomd
from hoomd import md
hoomd.context.initialize('--mode=cpu')
system = hoomd.init.read_gsd('file.gsd')
# groupB = hoomd.group.type(name="b-particles", type='B')
import freud
import gsd.hoomd
import numpy as np
import matplotlib.pyplot as plt
import hoomd
from hoomd import md
hoomd.context.initialize('--mode=cpu')
system = hoomd.init.read_gsd('file.gsd')
# groupB = hoomd.group.type(name="b-particles", type='B')
with gsd.hoomd.open(name='file.gsd', mode='rb') as t: frame = t.read_frame(0)
frame.particles.position[:].shape
frame.particles.typeid[:] == 0
frame.particles.typeid[:] == 0
frame.particles.position[frame.particles.typeid[:] == 0].shape
groupB = []
for pos in frame.particles.position[frame.particles.typeid == frame.particles.types.index('B')]: groupB.append(frame.particles.types.index('B'))
print(groupB)
nl = hoomd.md.nlist.tree()
# NVT integration
hoomd.md.integrate.mode_standard (dt = 0.005)
nvt = hoomd.md.integrate.nvt (group = groupB, kT = 1.2, tau = 1.0)
nvt.randomize_velocities (seed = 1)
# fslj = pair.force_shifted_lj(r_cut=453, nlist=nl)
# fslj.pair_coeff.set('B', 'B', epsilon=150.0, sigma=1.0)
pppm = hoomd.md.charge.pppm(groupB, nl)
pppm.set_params(Nx=150, Ny=150, Nz=250, order=6, rcut=450.0, alpha=0.007320808910601237)
# run the simulation
hoomd.run (100)
rdf = freud.density.RDF(bins=150, r_max=450)
def calc_rdf(timestep):
hoomd.util.quiet_status()
snap = system.take_snapshot()
hoomd.util.unquiet_status()
rdf.compute(system=snap, reset=False)
for pos in frame.particles.position[frame.particles.typeid == frame.particles.types.index('B')]: groupB.append(frame.particles.types.index('B'))
print(groupB)
nl = hoomd.md.nlist.tree()
# NVT integration
hoomd.md.integrate.mode_standard (dt = 0.005)
nvt = hoomd.md.integrate.nvt (group = groupB, kT = 1.2, tau = 1.0)
nvt.randomize_velocities (seed = 1)
# fslj = pair.force_shifted_lj(r_cut=453, nlist=nl)
# fslj.pair_coeff.set('B', 'B', epsilon=150.0, sigma=1.0)
pppm = hoomd.md.charge.pppm(groupB, nl)
pppm.set_params(Nx=150, Ny=150, Nz=250, order=6, rcut=450.0, alpha=0.007320808910601237)
# run the simulation
hoomd.run (100)
rdf = freud.density.RDF(bins=150, r_max=450)
def calc_rdf(timestep):
hoomd.util.quiet_status()
snap = system.take_snapshot()
hoomd.util.unquiet_status()
rdf.compute(system=snap, reset=False)
# Equilibrate the system a bit before accumulating the RDF.
hoomd.run(100)
hoomd.analyze.callback(calc_rdf, period=10)
hoomd.run(100)
# Store the computed RDF in a file
np.savetxt('rdf_273_100s_bins=150_period=10_N(x,y,z)=256_order=2.csv', np.vstack((rdf.bin_centers, rdf.rdf)).T,
delimiter=',', header='r, g(r)')
rdf_data = np.genfromtxt('rdf_273_100s_bins=150_period=10_N(x,y,z)=256_order=2.csv', delimiter=',')
plt.plot(rdf_data[:, 0], rdf_data[:, 1])
plt.title('Radial Distribution Function')
plt.xlabel('$r$')
plt.ylabel('$g(r)$')
plt.show()
Error message:
hoomd.run(100)
hoomd.analyze.callback(calc_rdf, period=10)
hoomd.run(100)
# Store the computed RDF in a file
np.savetxt('rdf_273_100s_bins=150_period=10_N(x,y,z)=256_order=2.csv', np.vstack((rdf.bin_centers, rdf.rdf)).T,
delimiter=',', header='r, g(r)')
rdf_data = np.genfromtxt('rdf_273_100s_bins=150_period=10_N(x,y,z)=256_order=2.csv', delimiter=',')
plt.plot(rdf_data[:, 0], rdf_data[:, 1])
plt.title('Radial Distribution Function')
plt.xlabel('$r$')
plt.ylabel('$g(r)$')
plt.show()
Error message:
----
Traceback (most recent call last):
File "/home/elsaid/untitled4.py", line 28, in <module>
nvt = hoomd.md.integrate.nvt (group = groupB, kT = 1.2, tau = 1.0)
File "/home/elsaid/anaconda3/lib/python3.8/site-packages/hoomd/md/integrate.py", line 221, in __init__
thermo = hoomd.compute._get_unique_thermo(group=group);
File "/home/elsaid/anaconda3/lib/python3.8/site-packages/hoomd/compute.py", line 282, in _get_unique_thermo
res = thermo(group);
File "/home/elsaid/anaconda3/lib/python3.8/site-packages/hoomd/compute.py", line 209, in __init__
if group.name != 'all':
AttributeError: 'list' object has no attribute 'name'
Traceback (most recent call last):
File "/home/elsaid/untitled4.py", line 28, in <module>
nvt = hoomd.md.integrate.nvt (group = groupB, kT = 1.2, tau = 1.0)
File "/home/elsaid/anaconda3/lib/python3.8/site-packages/hoomd/md/integrate.py", line 221, in __init__
thermo = hoomd.compute._get_unique_thermo(group=group);
File "/home/elsaid/anaconda3/lib/python3.8/site-packages/hoomd/compute.py", line 282, in _get_unique_thermo
res = thermo(group);
File "/home/elsaid/anaconda3/lib/python3.8/site-packages/hoomd/compute.py", line 209, in __init__
if group.name != 'all':
AttributeError: 'list' object has no attribute 'name'
Best,
Elsaid
To view this discussion on the web visit https://groups.google.com/d/msgid/hoomd-users/62199AD5-E778-4817-9451-418FF4576D8D%40gmail.com.
Eric Irrgang
Aug 17, 2021, 7:31:52 AM8/17/21
to hoomd...@googlegroups.com
Hi All,
Please note that the code snippets in my previous email were not intended as solutions to any particular problem. I only intended to illustrate possibilities.
I apologize for the formatting of the code snippets. I was using three '>' marks to indicate the Python interpreter prompt but that is common email syntax for quoting.
Given the context,
* `frame.particles.position[:].shape` would be expected to produce `(2961, 3)`
* `frame.particles.typeid[:] == 0` would produce a NumPy array boolean values with the same shape as the original
* `frame.particles.position[frame.particles.typeid[:] == 0].shape` would give `(2940, 3)`
I apologize for any confusion.
Please note that the code snippets in my previous email were not intended as solutions to any particular problem. I only intended to illustrate possibilities.
I apologize for the formatting of the code snippets. I was using three '>' marks to indicate the Python interpreter prompt but that is common email syntax for quoting.
Given the context,
* `frame.particles.position[:].shape` would be expected to produce `(2961, 3)`
* `frame.particles.typeid[:] == 0` would produce a NumPy array boolean values with the same shape as the original
* `frame.particles.position[frame.particles.typeid[:] == 0].shape` would give `(2940, 3)`
I apologize for any confusion.
Eric Irrgang
Aug 17, 2021, 7:39:39 AM8/17/21
to hoomd...@googlegroups.com
Hi Elsaid.
You are providing the wrong type of argument to hoomd.md.integrate.nvt.
https://hoomd-blue.readthedocs.io/en/stable/module-md-integrate.html#hoomd.md.integrate.nvt says that the `group` parameter is a `hoomd.group` object. You have provided a list.
To create a `hoomd.group` based on particle type, see https://hoomd-blue.readthedocs.io/en/stable/module-hoomd-group.html#hoomd.group.type
Note that creating a `hoomd.group` does not give the user any access to particles. It is only used in the parameters of other hoomd commands.
Best,
Eric
> To view this discussion on the web visit https://groups.google.com/d/msgid/hoomd-users/CAO9cOeBPHjhzEPn-G38kgw0gO2BumYURG4gT-sg4H%2BwFQdt%2B3w%40mail.gmail.com.
You are providing the wrong type of argument to hoomd.md.integrate.nvt.
https://hoomd-blue.readthedocs.io/en/stable/module-md-integrate.html#hoomd.md.integrate.nvt says that the `group` parameter is a `hoomd.group` object. You have provided a list.
To create a `hoomd.group` based on particle type, see https://hoomd-blue.readthedocs.io/en/stable/module-hoomd-group.html#hoomd.group.type
Note that creating a `hoomd.group` does not give the user any access to particles. It is only used in the parameters of other hoomd commands.
Best,
Eric
Elsaid Younes
Aug 25, 2021, 2:49:12 PM8/25/21
to hoomd...@googlegroups.com
Dear Eric,
I still cannot access type 'B'
hoomd.context.initialize('--mode=cpu')
system = hoomd.init.read_gsd('file.gsd')
with gsd.hoomd.open(name='file.gsd', mode='rb') as t: frame = t.read_frame(0)
frame.particles.position[:].shape
frame.particles.typeid[:] == 0
frame.particles.position[frame.particles.typeid[:] == 0].shape
groupB = []
for pos in frame.particles.position[frame.particles.typeid == frame.particles.types.index('B')]: groupB.append(frame.particles.types.index('B'))
print(groupB)
nl = hoomd.md.nlist.tree()
# NVT integration
frame.particles.position[:].shape
frame.particles.typeid[:] == 0
frame.particles.position[frame.particles.typeid[:] == 0].shape
groupB = []
for pos in frame.particles.position[frame.particles.typeid == frame.particles.types.index('B')]: groupB.append(frame.particles.types.index('B'))
print(groupB)
nl = hoomd.md.nlist.tree()
# NVT integration
groupB = hoomd.group.type('B');
hoomd.md.integrate.mode_standard (dt = 0.005)
nvt = hoomd.md.integrate.nvt (group = groupB, kT = 4.11433*(10**-24), tau = 1.0)
nvt.randomize_velocities (seed = 1)
hoomd.md.integrate.mode_standard (dt = 0.005)
nvt = hoomd.md.integrate.nvt (group = groupB, kT = 4.11433*(10**-24), tau = 1.0)
nvt.randomize_velocities (seed = 1)
pppm = hoomd.md.charge.pppm(groupB, nl)
pppm.set_params(Nx=150, Ny=150, Nz=250, order=6, rcut=450.0, alpha=0.007320808910601237)
# run the simulation
hoomd.run (100)
rdf = freud.density.RDF(bins=150, r_max=450)
def calc_rdf(timestep):
hoomd.util.quiet_status()
snap = system.take_snapshot()
hoomd.util.unquiet_status()
rdf.compute(system=snap, reset=False)
# Equilibrate the system a bit before accumulating the RDF.
hoomd.run(100)
hoomd.analyze.callback(calc_rdf, period=10)
hoomd.run(100)
# Store the computed RDF in a file
np.savetxt('rdf_273_100s_bins=150_period=10_N(x,y,z)=256_order=2.csv', np.vstack((rdf.bin_centers, rdf.rdf)).T,
delimiter=',', header='r, g(r)')
rdf_data = np.genfromtxt('rdf_273_100s_bins=150_period=10_N(x,y,z)=256_order=2.csv', delimiter=',')
plt.plot(rdf_data[:, 0], rdf_data[:, 1])
plt.title('Radial Distribution Function')
plt.xlabel('$r$')
plt.ylabel('$g(r)$')
plt.show()
When I don't use hoomd.group.type(), it gives me errors like :
File "/home/elsaid/untitled4.py", line 28, in <module>
nvt = hoomd.md.integrate.nvt (group = groupB, kT = 4.11433*(10**-24), tau = 1.0)
File "/home/elsaid/anaconda3/lib/python3.8/site-packages/hoomd/md/integrate.py", line 221, in __init__
thermo = hoomd.compute._get_unique_thermo(group=group);
File "/home/elsaid/anaconda3/lib/python3.8/site-packages/hoomd/compute.py", line 282, in _get_unique_thermo
res = thermo(group);
File "/home/elsaid/anaconda3/lib/python3.8/site-packages/hoomd/compute.py", line 209, in __init__
if group.name != 'all':
AttributeError: 'list' object has no attribute 'name'
Best,
Elsaid
To view this discussion on the web visit https://groups.google.com/d/msgid/hoomd-users/544ECB06-7F4D-45A3-B53B-E9C08186BB3A%40gmail.com.
Eric Irrgang
Aug 26, 2021, 10:36:17 AM8/26/21
to hoomd...@googlegroups.com
Hi Elsaid.
> When I don't use hoomd.group.type(), it gives me errors like :
>
> File "/home/elsaid/untitled4.py", line 28, in <module>
> nvt = hoomd.md.integrate.nvt (group = groupB, kT = 4.11433*(10**-24), tau = 1.0)
>
> File "/home/elsaid/anaconda3/lib/python3.8/site-packages/hoomd/md/integrate.py", line 221, in __init__
> thermo = hoomd.compute._get_unique_thermo(group=group);
>
> File "/home/elsaid/anaconda3/lib/python3.8/site-packages/hoomd/compute.py", line 282, in _get_unique_thermo
> res = thermo(group);
>
> File "/home/elsaid/anaconda3/lib/python3.8/site-packages/hoomd/compute.py", line 209, in __init__
> if group.name != 'all':
>
> AttributeError: 'list' object has no attribute 'name'
This is because you have assigned a list to `groupB` in the code above `groupB = hoomd.group.type('B');`, and a list is an invalid argument type for `hoomd.md.integrate.nvt (group = ...)`
> I still cannot access type 'B'
What do you mean by "access"?
Last time, I thought you meant that you wanted to inspect the particle data for particles of type 'B', so I suggested some syntax for exploring the particle data structure. But then it seemed like you were trying to define the group to be passed to the integrator, so we suggested `hoomd.group.type()`.
At a quick glance, your script looks fine. Are you getting an error?
> hoomd.md.integrate.mode_standard (dt = 0.005)
> nvt = hoomd.md.integrate.nvt (group = groupB, kT = 4.11433*(10**-24), tau = 1.0)
Depending on your other basic units, this temperature may be approximately equal to absolute zero. Also, if you are using the single-precision build (default), the equations of motion may evaluate to zero change due to floating point precision issues.
It's generally a good idea to choose units in HOOMD such that important values are as close to 1 in magnitude as possible. More than three orders of magnitude above or below 1 for parameters can lead to precision problems (too much truncation error).
See also https://hoomd-blue.readthedocs.io/en/stable/units.html
Best,
Eric
> When I don't use hoomd.group.type(), it gives me errors like :
>
> File "/home/elsaid/untitled4.py", line 28, in <module>
> nvt = hoomd.md.integrate.nvt (group = groupB, kT = 4.11433*(10**-24), tau = 1.0)
>
> File "/home/elsaid/anaconda3/lib/python3.8/site-packages/hoomd/md/integrate.py", line 221, in __init__
> thermo = hoomd.compute._get_unique_thermo(group=group);
>
> File "/home/elsaid/anaconda3/lib/python3.8/site-packages/hoomd/compute.py", line 282, in _get_unique_thermo
> res = thermo(group);
>
> File "/home/elsaid/anaconda3/lib/python3.8/site-packages/hoomd/compute.py", line 209, in __init__
> if group.name != 'all':
>
> AttributeError: 'list' object has no attribute 'name'
> I still cannot access type 'B'
Last time, I thought you meant that you wanted to inspect the particle data for particles of type 'B', so I suggested some syntax for exploring the particle data structure. But then it seemed like you were trying to define the group to be passed to the integrator, so we suggested `hoomd.group.type()`.
At a quick glance, your script looks fine. Are you getting an error?
> hoomd.md.integrate.mode_standard (dt = 0.005)
> nvt = hoomd.md.integrate.nvt (group = groupB, kT = 4.11433*(10**-24), tau = 1.0)
It's generally a good idea to choose units in HOOMD such that important values are as close to 1 in magnitude as possible. More than three orders of magnitude above or below 1 for parameters can lead to precision problems (too much truncation error).
See also https://hoomd-blue.readthedocs.io/en/stable/units.html
Best,
Eric
Joshua Anderson
Aug 26, 2021, 1:09:22 PM8/26/21
to hoomd...@googlegroups.com
All,
I just want to clarify that single precision is not the default. All HOOMD binaries I provide are double precision, as is the default setting when you build from source.
------
Joshua A. Anderson, Ph.D.
Research Area Specialist, Chemical Engineering, University of Michigan
I just want to clarify that single precision is not the default. All HOOMD binaries I provide are double precision, as is the default setting when you build from source.
------
Joshua A. Anderson, Ph.D.
Research Area Specialist, Chemical Engineering, University of Michigan
> --
> You received this message because you are subscribed to the Google Groups "hoomd-users" group.
> To unsubscribe from this group and stop receiving emails from it, send an email to hoomd-users...@googlegroups.com.
> To view this discussion on the web visit https://groups.google.com/d/msgid/hoomd-users/D7DC66BC-5F31-4C14-BA2C-9E4B7D50B6CA%40gmail.com.
> You received this message because you are subscribed to the Google Groups "hoomd-users" group.
> To unsubscribe from this group and stop receiving emails from it, send an email to hoomd-users...@googlegroups.com.
Elsaid Younes
Aug 26, 2021, 2:21:54 PM8/26/21
to hoomd...@googlegroups.com
Dear Eric,
since I have two types of particles ['A','B'] with different diameters)) here are the calculated r.d.f.When I write groupB = hoomd.group.type('B'), it does not pass that type to the NVT, the r.d.f. calculated for the other type ((
For hoomd.group.type('A'):
HOOMD-blue 2.9.6 DOUBLE HPMC_MIXED TBB SSE SSE2 SSE3
Compiled: 03/17/2021
Copyright (c) 2009-2019 The Regents of the University of Michigan.
-----
You are using HOOMD-blue. Please cite the following:
* J A Anderson, J Glaser, and S C Glotzer. "HOOMD-blue: A Python package for
high-performance molecular dynamics and hard particle Monte Carlo
simulations", Computational Materials Science 173 (2020) 109363
-----
HOOMD-blue is running on the CPU
notice(2): Group "all" created containing 2961 particles
-----
You are using tree neighbor lists. Please cite the following:
* M P Howard, J A Anderson, A Nikoubashman, S C Glotzer, and A Z
Panagiotopoulos. "Efficient neighbor list calculation for molecular simulation
of colloidal systems using graphics processing units", Computer Physics
Communications 203 (2016) 45--52
* M P Howard, A Statt, F Madutsa, T M Truskett, and A Z Panagiotopoulos.
"Quantized bounding volume hierarchies for neighbor search in molecular
simulations on graphics processing units", Computational Materials Science 164
(2019) 139--146
-----
notice(2): Group "type A" created containing 2940 particles
-----
You are using PPPM. Please cite the following:
* D N LeBard, B G Levine, S A Barr, A Jusufi, S Sanders, M L Klein, and A Z
Panagiotopoulos. "Self-assembly of coarse-grained ionic surfactants
accelerated by graphics processing units", Journal of Computational Physics 8
(2012) 2385-2397
-----
notice(2): -- Neighborlist exclusion statistics -- :
notice(2): Particles with 0 exclusions : 2961
notice(2): Neighbors included by diameter : no
notice(2): Neighbors excluded when in the same body: no
** starting run **
notice(2): charge.pppm: RMS error: 5.39052e-14
Compiled: 03/17/2021
Copyright (c) 2009-2019 The Regents of the University of Michigan.
-----
You are using HOOMD-blue. Please cite the following:
* J A Anderson, J Glaser, and S C Glotzer. "HOOMD-blue: A Python package for
high-performance molecular dynamics and hard particle Monte Carlo
simulations", Computational Materials Science 173 (2020) 109363
-----
HOOMD-blue is running on the CPU
notice(2): Group "all" created containing 2961 particles
-----
You are using tree neighbor lists. Please cite the following:
* M P Howard, J A Anderson, A Nikoubashman, S C Glotzer, and A Z
Panagiotopoulos. "Efficient neighbor list calculation for molecular simulation
of colloidal systems using graphics processing units", Computer Physics
Communications 203 (2016) 45--52
* M P Howard, A Statt, F Madutsa, T M Truskett, and A Z Panagiotopoulos.
"Quantized bounding volume hierarchies for neighbor search in molecular
simulations on graphics processing units", Computational Materials Science 164
(2019) 139--146
-----
notice(2): Group "type A" created containing 2940 particles
-----
You are using PPPM. Please cite the following:
* D N LeBard, B G Levine, S A Barr, A Jusufi, S Sanders, M L Klein, and A Z
Panagiotopoulos. "Self-assembly of coarse-grained ionic surfactants
accelerated by graphics processing units", Journal of Computational Physics 8
(2012) 2385-2397
-----
notice(2): -- Neighborlist exclusion statistics -- :
notice(2): Particles with 0 exclusions : 2961
notice(2): Neighbors included by diameter : no
notice(2): Neighbors excluded when in the same body: no
** starting run **
notice(2): charge.pppm: RMS error: 5.39052e-14
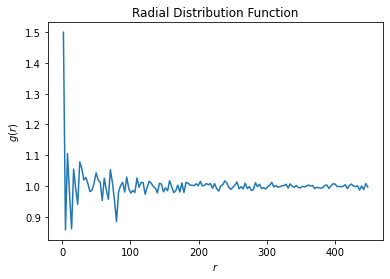
For hoomd.group.type('B'):
--
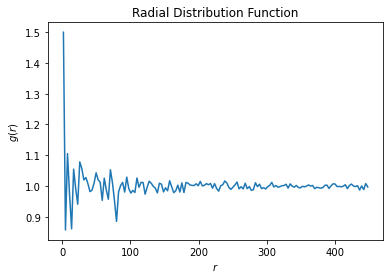
notice(2): Group "all" created containing 2961 particles
-----
You are using tree neighbor lists. Please cite the following:
* M P Howard, J A Anderson, A Nikoubashman, S C Glotzer, and A Z
Panagiotopoulos. "Efficient neighbor list calculation for molecular simulation
of colloidal systems using graphics processing units", Computer Physics
Communications 203 (2016) 45--52
* M P Howard, A Statt, F Madutsa, T M Truskett, and A Z Panagiotopoulos.
"Quantized bounding volume hierarchies for neighbor search in molecular
simulations on graphics processing units", Computational Materials Science 164
(2019) 139--146
-----
notice(2): Group "type B" created containing 21 particles
-----
You are using PPPM. Please cite the following:
* D N LeBard, B G Levine, S A Barr, A Jusufi, S Sanders, M L Klein, and A Z
Panagiotopoulos. "Self-assembly of coarse-grained ionic surfactants
accelerated by graphics processing units", Journal of Computational Physics 8
(2012) 2385-2397
-----
notice(2): -- Neighborlist exclusion statistics -- :
notice(2): Particles with 0 exclusions : 2961
notice(2): Neighbors included by diameter : no
notice(2): Neighbors excluded when in the same body: no
** starting run **
notice(2): charge.pppm: RMS error: 7.54673e-12
-----
You are using tree neighbor lists. Please cite the following:
* M P Howard, J A Anderson, A Nikoubashman, S C Glotzer, and A Z
Panagiotopoulos. "Efficient neighbor list calculation for molecular simulation
of colloidal systems using graphics processing units", Computer Physics
Communications 203 (2016) 45--52
* M P Howard, A Statt, F Madutsa, T M Truskett, and A Z Panagiotopoulos.
"Quantized bounding volume hierarchies for neighbor search in molecular
simulations on graphics processing units", Computational Materials Science 164
(2019) 139--146
-----
notice(2): Group "type B" created containing 21 particles
-----
You are using PPPM. Please cite the following:
* D N LeBard, B G Levine, S A Barr, A Jusufi, S Sanders, M L Klein, and A Z
Panagiotopoulos. "Self-assembly of coarse-grained ionic surfactants
accelerated by graphics processing units", Journal of Computational Physics 8
(2012) 2385-2397
-----
notice(2): -- Neighborlist exclusion statistics -- :
notice(2): Particles with 0 exclusions : 2961
notice(2): Neighbors included by diameter : no
notice(2): Neighbors excluded when in the same body: no
** starting run **
notice(2): charge.pppm: RMS error: 7.54673e-12

For units:
I am very confused. How shall I write:
"columb"? charge = -140 columb
"Temperature"? T = 298 K, boltzmann constant* T = kT = 4.11433*(10**-24)
Best,.
Elsaid
--
You received this message because you are subscribed to the Google Groups "hoomd-users" group.
To unsubscribe from this group and stop receiving emails from it, send an email to hoomd-users...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/hoomd-users/D7DC66BC-5F31-4C14-BA2C-9E4B7D50B6CA%40gmail.com.
Joshua Anderson
Aug 26, 2021, 2:28:47 PM8/26/21
to hoomd...@googlegroups.com
Elsaid,
The group that you pass to the NVT integration method sets which particle that method acts on. It DOES NOT set what particles freud uses when computing RDFs.
> To view this discussion on the web visit https://groups.google.com/d/msgid/hoomd-users/CAO9cOeAq3fCng8GcVON%3Du85t2%3DLg9Uz5%2BT_uGux58RXEM7goqA%40mail.gmail.com.
The group that you pass to the NVT integration method sets which particle that method acts on. It DOES NOT set what particles freud uses when computing RDFs.
------
Joshua A. Anderson, Ph.D.
Research Area Specialist, Chemical Engineering, University of Michigan
Joshua A. Anderson, Ph.D.
Research Area Specialist, Chemical Engineering, University of Michigan
Eric Irrgang
Aug 27, 2021, 6:20:18 AM8/27/21
to hoomd...@googlegroups.com
Following on Josh's note, it doesn't look like you are applying any integrator to particles of type 'A'. This is fine if you intend for them to be constrained or if they are part of a rigid body, but I thought I should point that out.
P.S. Thanks to Josh for correcting me on my mistake on default precision.
P.S. Thanks to Josh for correcting me on my mistake on default precision.
Eric Irrgang
Aug 27, 2021, 6:55:54 AM8/27/21
to hoomd...@googlegroups.com
> For units:
> I am very confused. How shall I write:
> "columb"? charge = -140 columb
> "Temperature"? T = 298 K, boltzmann constant* T = kT = 4.11433*(10**-24)
Regardless of floating-point precision, it is usually convenient to choose at least a few units such that many quantities are 1.0 (whether that is particle mass, charge, size, temperature...).
You might get more feedback from HOOMD users doing electrostatics simulations if you start a new thread to ask what combinations of units people find most useful for simulations like yours (similar temperature, particle size, charge magnitude, etc)
Best,
Eric
Reply all
Reply to author
Forward
0 new messages
