Communication problems between DSI and Brainstorm
123 views
Skip to first unread message
Margherita Matarrese
May 5, 2021, 9:14:52 AM5/5/21
to DSI Studio
Dear DSI users,
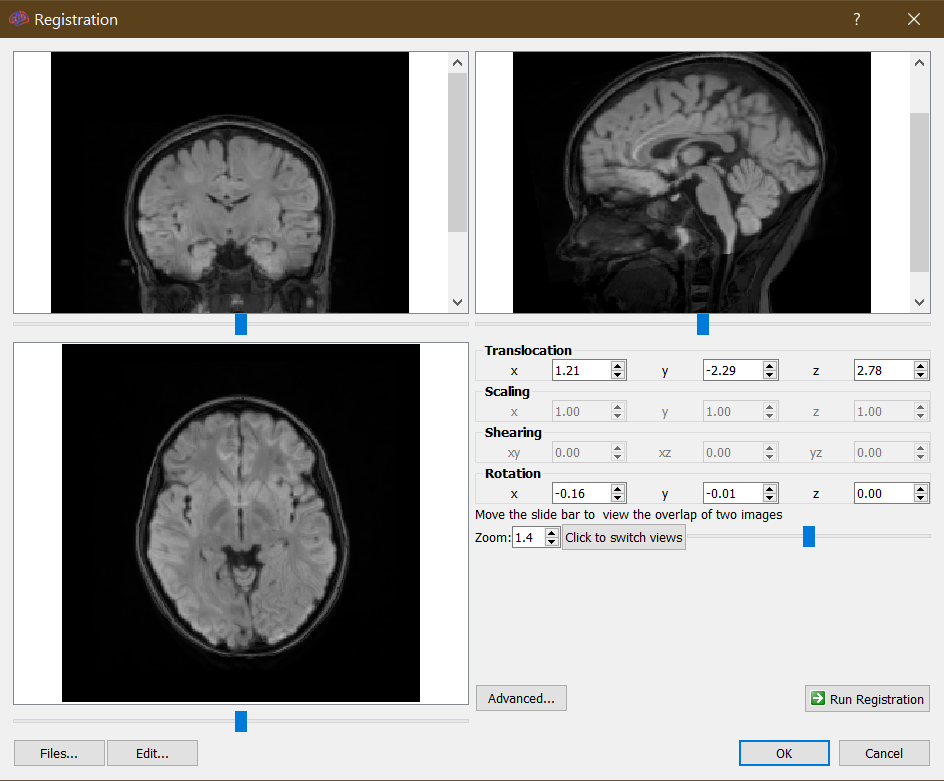
I would like to submit a problem that I am struggling with these days hoping to have helpful suggestions from you. I am trying to standardize the data exported from DSI with Brainstorm (https://neuroimage.usc.edu/brainstorm) convention.
What I am trying to find is the affine transformation that allows me to visualize the fibers in a specific voxel space. Brainstorm uses a specific subject coordinate system (SCS) but the affine transformation from MRI to SCS coordinates is fully described with a rotation matrix [3x3] and a translation matrix [3x1].
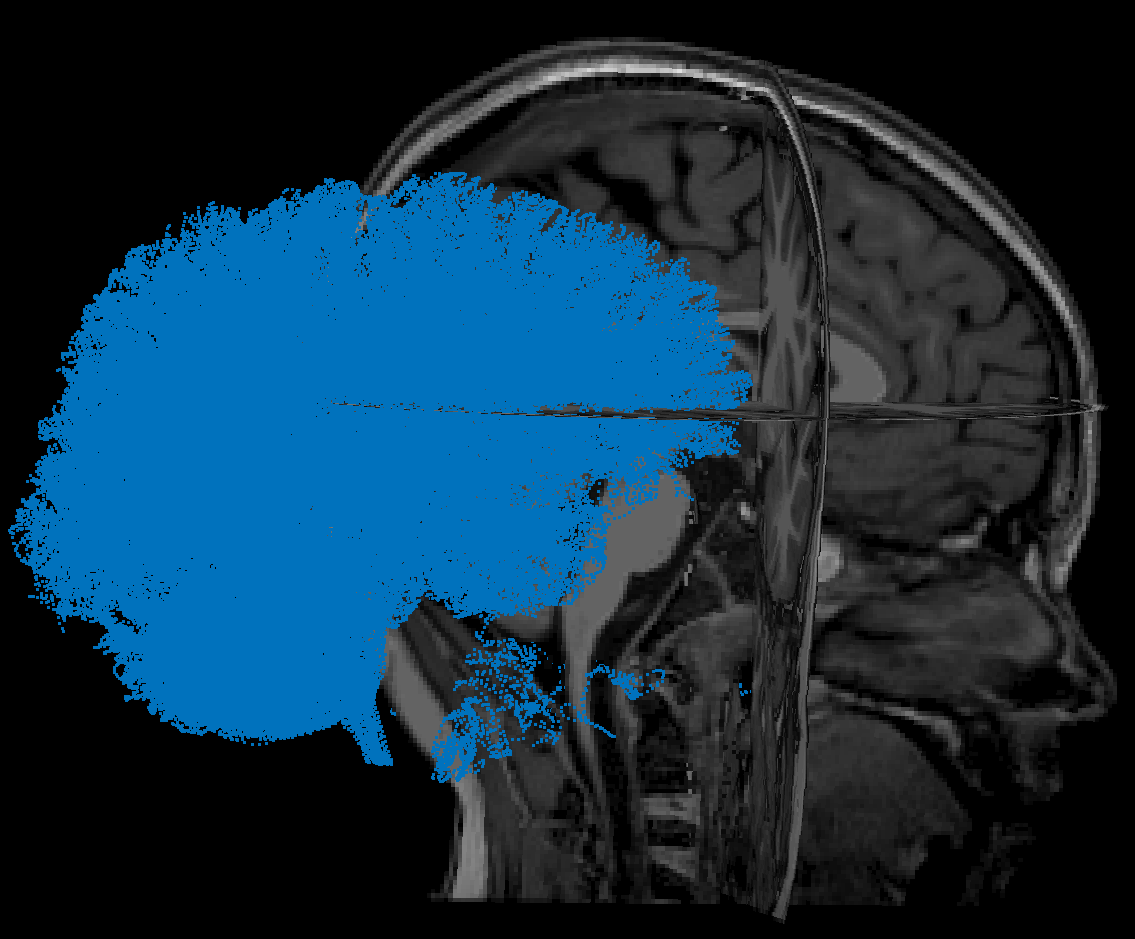
In DSI, I have used T1 MRI which uses Braistorm to work. But what I can't understand is how to align the DTI information to the same voxel space as the MRI. When I try to do the alignment in DSI between the two images, in the menu I see that there is actually a translocation and a rotation. Would it be possible to define, or rather export, a rotation and/or roto-translation matrix?

In fact, by exporting the fibers and applying the transformation in the Brainstorm space, I can get comparable information but the initial roto-translation between MRI and DTI is evidently not maintained for the fibers.
Could any of you help me get the right coordinate transformation? Or at least how to align all the DCM files with the T1? I tried with the "O6 tool" but I don't understand two things:
- how to load all the dcm files (loading more than one file, the program crashes);
- how to export the actually rotated dcm files.
Thank you very much for your help,
Margherita
Frank Yeh
May 5, 2021, 9:28:13 AM5/5/21
to dsi-s...@googlegroups.com
Dealing with the transformation matrix directly will be very challenging because different tools have their own way of using the rotation and translation matrix.
If your purpose is to output tracts to the T1W voxel coordinates, then [Tracts][Save Tracts][Save Current Tracts in Slice (T1W/T2W) Space] can help bypass the problem.
On Wed, May 5, 2021 at 9:14 AM Margherita Matarrese <mita...@gmail.com> wrote:
Could any of you help me get the right coordinate transformation? Or at least how to align all the DCM files with the T1? I tried with the "O6 tool" but I don't understand two things:
- how to load all the dcm files (loading more than one file, the program crashes);
Sorry the O6 Tool has not yet support multiple DCM (I will revise DSI Studio to enable this support). For the time being, you can convert DCM to NIFTI files using dcm2nii.
- how to export the actually rotated dcm files.
Writing to DCM files is not currently possible in DSI Studio because of the complexity of the DICOM header.
DSI Studio can output NIFTI file and you may convert NIFTI to DCM using Matlab or other tools.
Hope this helps.
Frank
Thank you very much for your help,Margherita
--
You received this message because you are subscribed to the Google Groups "DSI Studio" group.
To unsubscribe from this group and stop receiving emails from it, send an email to dsi-studio+...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/dsi-studio/173846fc-074b-4e32-aaac-659552de0ac1n%40googlegroups.com.
Margherita Matarrese
May 18, 2021, 4:31:37 AM5/18/21
to DSI Studio
Thank you very much Frank for your reply. After several attempts, I managed to find the correct transformation. If it might interest you we can discuss it better together.
I also wanted to ask you if you could provide me with some bibliographical material on the three fiber tracking algorithms currently implemented in DSI studio. I noticed that in my specific application only voxel tracking seems to perform well and I was wondering why.
Thanks again,
Margherita
Frank Yeh
May 18, 2021, 10:22:31 AM5/18/21
to dsi-s...@googlegroups.com
The first two tracking methods are conventional deterministic fiber trackings: https://pubmed.ncbi.nlm.nih.gov/24348913/
The "voxel tracking" one does not have a publication yet. It is more of a voxel-wise clustering method that connects neighboring voxels that form a connecting trajectory.
Frank
To view this discussion on the web visit https://groups.google.com/d/msgid/dsi-studio/2152d8eb-4f67-488e-a9f0-12152b267801n%40googlegroups.com.
yi lei
Jul 24, 2022, 1:17:38 AM7/24/22
to DSI Studio
HI, could u please share the way how to correct the trk transformation?
Margherita Matarrese
Jul 24, 2022, 2:07:26 PM7/24/22
to DSI Studio
Dear Frank,
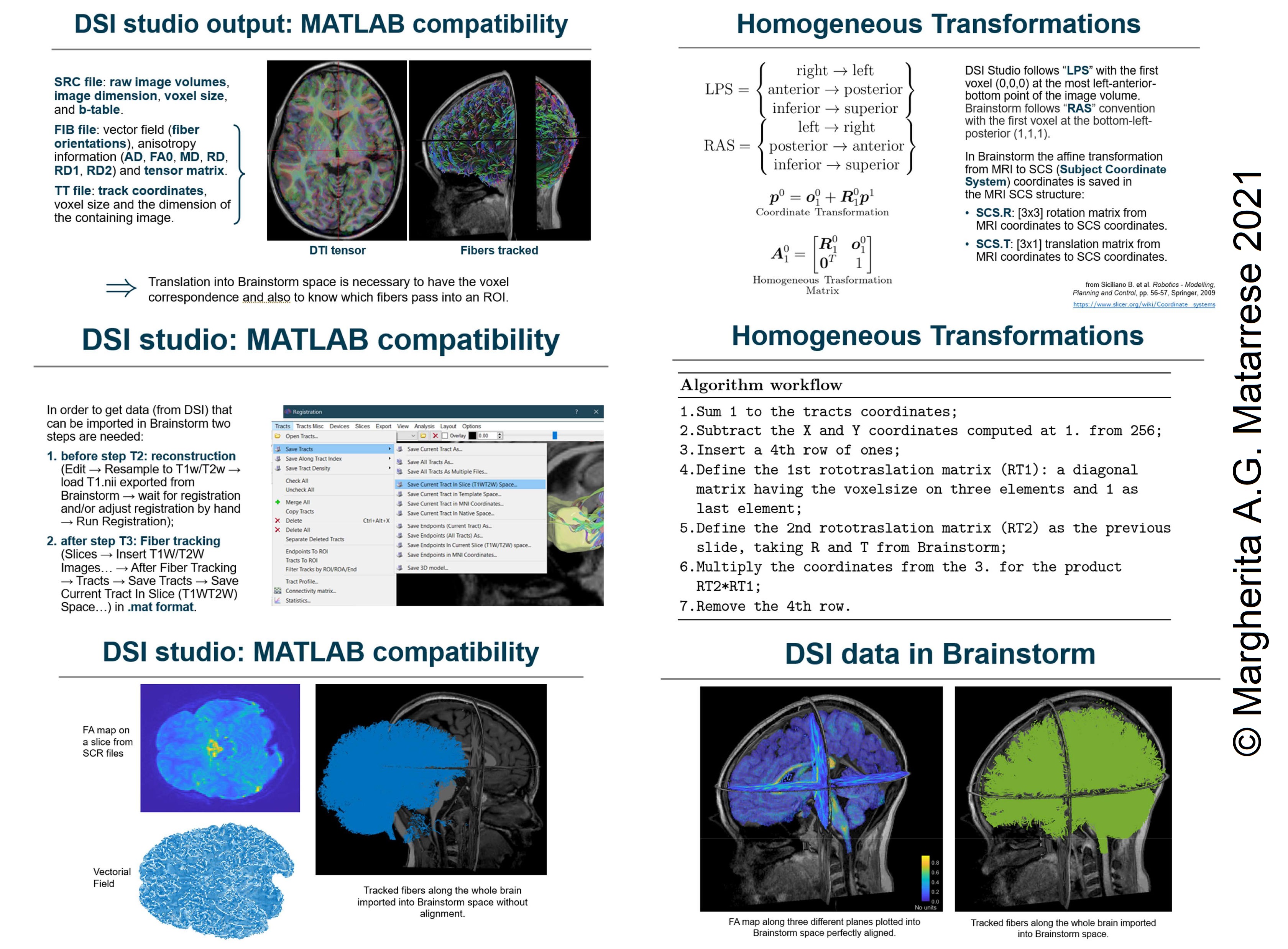
I found some slides I had done last year. Hope they can help. Let me know if I can help further.
Margherita

Fang-Cheng Yeh
Jul 24, 2022, 4:25:31 PM7/24/22
to dsi-s...@googlegroups.com
Thanks! This is very helpful!
Frank
To view this discussion on the web visit https://groups.google.com/d/msgid/dsi-studio/71dec281-6aa5-4ab2-b890-40ca9af51a9dn%40googlegroups.com.
Reply all
Reply to author
Forward
0 new messages
