Dht2 function and binned distaance
Delphine Ducros
Region.Label Area Effort distbegin distend Sample.Label
1 Nord 8961 4.09 NA NA 1
2 Nord 8961 3.87 NA NA 2
3 Nord 8961 39.63 0.08 0.18 3
4 Nord 8961 39.63 0.08 0.18 3
5 Nord 8961 39.63 0.08 0.18 3
6 Nord 8961 39.63 0.00 0.08 3dht2(ddf=hn.cos2.df, flatfile=data, strat_formula = ~ Region.Label, stratification = "geographical")
Warning messages:
1: In `[<-.data.frame`(`*tmp*`, flatfile$Sample.Label %in% sl_diff, :
provided 7 variables to replace 6 variables
2: In `[<-.data.frame`(`*tmp*`, flatfile$Sample.Label %in% sl_diff, :
provided 7 variables to replace 6 variables
Summary statistics:
Region.Label Area CoveredArea Effort n k ER se.ER cv.ER
2 22407 2327.068 1491.71 0 18 0 0 0
Nord 8961 4690.608 3006.80 0 87 0 0 0
Sud 10699 3586.924 2299.31 0 42 0 0 0
Total 42067 10604.599 6797.82 0 147 0 0 0
Abundance estimates:
Region.Label Estimate se cv LCI UCI df
2 0 0 0 0 0 1
Nord 0 0 0 0 0 1
Sud 0 0 0 0 0 1
Total 0 0 NaN NaN NaN NaN
Component percentages of variance:
Region.Label Detection ER
2 NaN NaN
Nord NaN NaN
Sud NaN NaN
Total 100 0
Eric Rexstad
Delphine
Thanks for your report about problems with `dht2`. We'll have a look at the problem and may contact you off-line about particulars.
Meanwhile, you should be able to work around the problem in the manner you described: replace the two columns distbegin and distend with a single column named "distance", into that column place the midpoint of the bin (e.g. below your end points are 0.08 and 0.18, use the value 0.13 for distance.
Then use the "cutpoints=" argument in your call to ds() to
conduct your binned analysis. You should be able to send the
result of that call to ds() off to dht2(). Let us know if that
works for you.
--
You received this message because you are subscribed to the Google Groups "distance-sampling" group.
To unsubscribe from this group and stop receiving emails from it, send an email to distance-sampl...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/distance-sampling/b73d3a81-b8f0-453a-a1b9-95501257ac83o%40googlegroups.com.
-- Eric Rexstad Centre for Ecological and Environmental Modelling University of St Andrews St Andrews is a charity registered in Scotland SC013532
David Lawrence Miller
Thanks for taking dht2 for a spin and sorry you had some problems. Would
you mind sending me your data off-list and I'll take a look at what's
going on here?
cheers,
--dave
On 23/07/2020 16:14, Delphine Ducros wrote:
> You received this message because you are subscribed to the Google
> Groups "distance-sampling" group.
> To unsubscribe from this group and stop receiving emails from it, send
> an email to distance-sampl...@googlegroups.com
> https://groups.google.com/d/msgid/distance-sampling/b73d3a81-b8f0-453a-a1b9-95501257ac83o%40googlegroups.com
DEFOS DU RAU Pierre
Hello & Thank you Eric and Delphine. But when it comes to multispecies DS survey with binned distance (e.g. songbird by point or seabirds by transect), would it make sense to estimate local guild species richness in a DS framework, as one can expect to detect all species in the closest bin but less so in the further bins? Would it be theoretically “allowed” ??? Of course, assuming models would converge, regional size estimates would make no sense but DSM application could be partially interesting by outputting a layer of local sp richness, no?
Sorry if I am totally misleading but curiosity nibbles me (as we say in French)
Thanks a lot for all the so fast replies to any questions almost everyday on this list!
Pierre
De : 'Eric Rexstad' via distance-sampling <distance...@googlegroups.com>
Envoyé : jeudi 23 juillet 2020 19:42
À : Delphine Ducros <ducros....@gmail.com>; distance-sampling <distance...@googlegroups.com>
Objet : Re: [distance-sampling] Dht2 function and binned distaance
To view this discussion on the web visit https://groups.google.com/d/msgid/distance-sampling/8728ba75-bfea-0261-0b03-d2c996723510%40st-andrews.ac.uk.
Delphine Ducros
To unsubscribe from this group and stop receiving emails from it, send an email to distance...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/distance-sampling/b73d3a81-b8f0-453a-a1b9-95501257ac83o%40googlegroups.com.
David Lawrence Miller
get to the bottom of this.
The warnings were caused by a bug in how pre-binned data was handled in
dht2 (i.e., when columns distend and distbegin were present but distance
is not). This is fixed in the latest versions of Distance on github and
will be included in the forthcoming CRAN release.
In the meantime the fix suggested by Eric seems to do the job if you
don't want to use the development version of the package.
cheers,
--dave
On 23/07/2020 16:14, Delphine Ducros wrote:
> You received this message because you are subscribed to the Google
> Groups "distance-sampling" group.
> To unsubscribe from this group and stop receiving emails from it, send
> <mailto:distance-sampl...@googlegroups.com>.
> https://groups.google.com/d/msgid/distance-sampling/b73d3a81-b8f0-453a-a1b9-95501257ac83o%40googlegroups.com
Delphine Ducros
> an email to distance...@googlegroups.com
> <mailto:distance-sampling+unsub...@googlegroups.com>.
Annika Zuleger

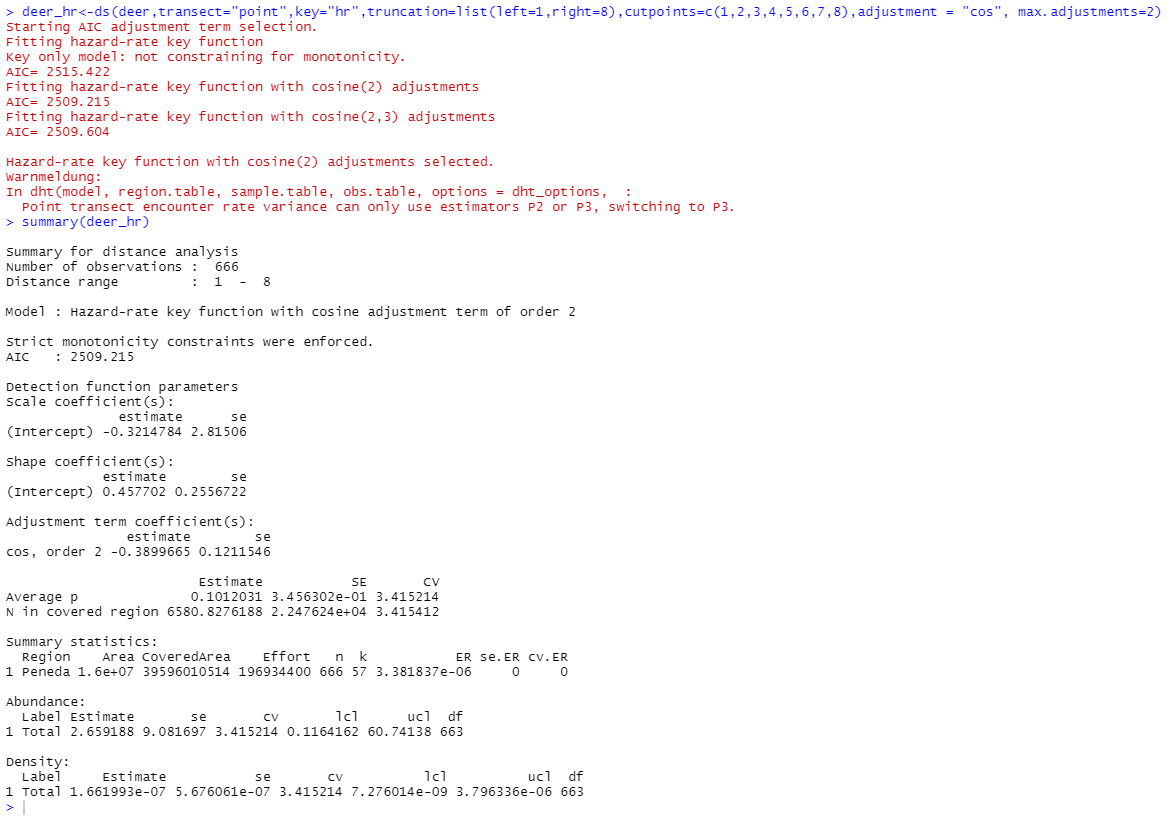
However, when I use the dht2 function I receive the same warning and incomplete output (see below, in this example my data also includes transects without observations, but the problems remains when I run the analysis only for those transects where the species was observed).

Eric Rexstad
Annika
A quick read of your question suggests the problem might be at the end of your email, rather than the beginning.
I think the problem you are having may have been fixed last summer (when the correspondence between Delphine and Dave Miller took place). That version of the Distance package 1.0.2 was loaded onto CRAN last December.
I suggest you install version 1.0.2 of
Distance from CRAN (which hopefully will circumvent the "fail to
install Distance from Github" problem). Try running your
analysis with that version of the package and see what happens.
If the problems persist, we will further investigate.
To unsubscribe from this group and stop receiving emails from it, send an email to distance-sampl...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/distance-sampling/5c21290f-7927-4695-b37d-f2d8e61f83c0n%40googlegroups.com.
Annika Zuleger
Eric Rexstad
Annika
OK, that eliminates the simple solution,
meaning we need to investigate more complex solutions. Would
you be willing to send your data set and the code you have run
off-list to me so I can investigate further?
To view this discussion on the web visit https://groups.google.com/d/msgid/distance-sampling/c1112c26-55ac-4a4a-a76f-ef739f512e80n%40googlegroups.com.
Annika Zuleger

