Unsure how to import Excel file with all columns
217 views
Skip to first unread message
Alfred Hargrave
Nov 1, 2020, 1:22:52 PM11/1/20
to cytoscape-helpdesk
Dear all,
I'm new to Cytoscape and I've been trying out the tutorials and reading the manual, but I can't find a solution to my problem.
I'm trying to create a PPI network from proteomics results with the color of the nodes as fold change in expression. When I paste the proteins (GeneName) into STRING, I get a network that looks great, but I want the additional features, so I'm copying a tutorial with my own data.
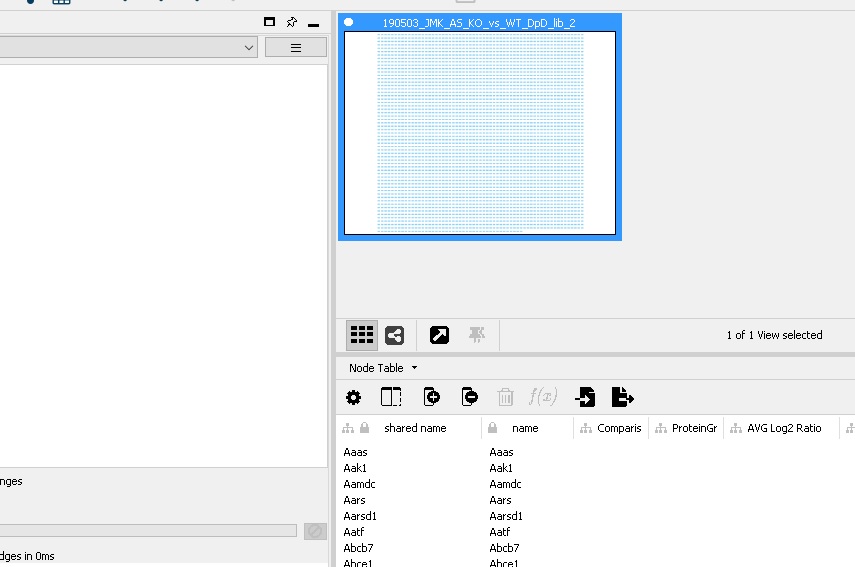
So first I imported Network from File with all columns switched off but "GeneName" set as source node. So far so good. Then when I try to import Table from File with the same file with the source node also set as "GeneName", I get the result shown below: a Node Table with no numerical columns in it.
Am I uploading with the wrong file type? (.xlsx, .tsv, .txt, .tsv?).
I'm also unsure how we set what the source node and target nodes as for the connections between nodes. In STRING this is done automatically, the network is built from a database.
Many thanks,
Alfred

Alex Pico
Nov 1, 2020, 1:46:01 PM11/1/20
to cytoscape-helpdesk
Hi Alfred,
Here’s how I think about data visualization in Cytoscape:
1. Network. Starting with STRING is a great way to do. Using the same names found in your dataset, i.e., the “GeneName” column will be helpful in the next step. The STRING network will load with its own node names, but there will be a column called “query term” that will match the names you used.
2. Data. After you have a network, Import Table from File to associate your data values with the nodes in the network. You will need to tell Cytoscape which column in the network Node Table (“query term”) matches which column in your dataset (“GeneName”). In whichever rows those columns match, your data values will be loaded.
3. Styles.Now open the Style panel to tell Cytoscape how you want particular columns of data (e.g., “fold change”) to map to visual properties (e.g., node fill color).
Let us know if you get stuck on any of these steps in particular and share screenshots and files so we can reproduce the steps and suggest solutions. I’m making some guesses about your use case in the above steps that may not be correct.
- Alex
<Untitled.jpg>--
You received this message because you are subscribed to the Google Groups "cytoscape-helpdesk" group.
To unsubscribe from this group and stop receiving emails from it, send an email to cytoscape-helpd...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/cytoscape-helpdesk/0dfb1991-45bf-4f48-b2ca-7e17d8171014n%40googlegroups.com.
<Untitled.jpg>
Alfred Hargrave
Nov 3, 2020, 2:02:05 PM11/3/20
to cytoscape...@googlegroups.com
Hi Alex,
" You will need to tell Cytoscape which column in the network Node Table (“query term”) matches which column in your dataset (“GeneName”). "
Do you mean that I set "Key Column for Network" to "query term" and set "GeneName" as the source node?
Many thanks,
Alfred
You received this message because you are subscribed to a topic in the Google Groups "cytoscape-helpdesk" group.
To unsubscribe from this topic, visit https://groups.google.com/d/topic/cytoscape-helpdesk/LA6s1S7fPg4/unsubscribe.
To unsubscribe from this group and all its topics, send an email to cytoscape-helpd...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/cytoscape-helpdesk/5D674FBB-38D0-4CA2-A4F9-17B7AB6C19FA%40gladstone.ucsf.edu.
Alex Pico
Nov 3, 2020, 3:21:25 PM11/3/20
to cytoscape...@googlegroups.com
Yes, after the STING network is loaded, the dialog for Import > Table from File… should have “Key Column for Network,” which you would set to “query term.” And the “Key” column in the “Preview” section should be designated for your “GeneName” column. It will be set to the first, left-most column by default, so you may or may not have to change it.
You should not see a choice for “source node” in this dialog. If you do see it, then you may have picked Import > Network from File… by mistake.
- Alex
To view this discussion on the web visit https://groups.google.com/d/msgid/cytoscape-helpdesk/CAGvYsho7LM9WOn9Jcb9%2BmiJxhJHfXNdCzvKrcy60pfk4eDCj8A%40mail.gmail.com.
Alfred Hargrave
Nov 7, 2020, 12:28:36 PM11/7/20
to cytoscape...@googlegroups.com
Thank you Alex, I've now successfully done that.
Do you have any recommendations on how to perform a functional enrichment on the clusters that I can see in my network? I clicked on clusters in the far right, which broke my network down into identifiable clusters, then I clicked on functional enrichment, but it said that none was found. Am I doing it correctly?
Many thanks,
Alfred
To view this discussion on the web visit https://groups.google.com/d/msgid/cytoscape-helpdesk/06A39CB2-B0F9-4AC8-8148-CAAA651BBD90%40gladstone.ucsf.edu.
Alex Pico
Nov 7, 2020, 2:49:29 PM11/7/20
to cytoscape-helpdesk
If you see a notice in the STRING Enrichment tab of the Node Table that says, “No enrichment has bee retrieved for this network,” then you’ll want to run the enrichment from the menu: Apps > STRING Enrichment > Retrieve functional enrichment.
If you have run the enrichment analysis (which involves a dialog with some options and a task that takes a couple seconds to run) and the results are still empty, then you have selected a set of nodes that do not have any enrichment results.
- Alex
Alfred Hargrave
Nov 11, 2020, 6:35:50 AM11/11/20
to cytoscape...@googlegroups.com
Thanks Alex that's really helped! Is there a quick way to identify the nodes in my network with the greatest number of arcs to other nodes?
--
You received this message because you are subscribed to a topic in the Google Groups "cytoscape-helpdesk" group.
To unsubscribe from this topic, visit https://groups.google.com/d/topic/cytoscape-helpdesk/LA6s1S7fPg4/unsubscribe.
To unsubscribe from this group and all its topics, send an email to cytoscape-helpd...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/cytoscape-helpdesk/8FD2EAAA-7F53-4A74-B998-D57D4D9D1FAA%40gladstone.ucsf.edu.
Alex Pico
Nov 11, 2020, 3:04:51 PM11/11/20
to cytoscape-helpdesk
This should give you a column of node degree (numbers edges) per node:
- Alex
You received this message because you are subscribed to the Google Groups "cytoscape-helpdesk" group.
To unsubscribe from this group and stop receiving emails from it, send an email to cytoscape-helpd...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/cytoscape-helpdesk/CAGvYshrDsq%2BH%2BZn5BSJxRYoJO4nt1HP4cab%2BktT_9gFB53%3D4fQ%40mail.gmail.com.
Alfred Hargrave
Nov 29, 2020, 9:01:06 AM11/29/20
to cytoscape...@googlegroups.com
Hi Alex,
When I try to find the node centrality by following the instructions in your link (Analyze network > as directed graph) I get the following error: NetworkInterpretation is null. Do you know how to resolve this?
Many thanks, Alfred.
To view this discussion on the web visit https://groups.google.com/d/msgid/cytoscape-helpdesk/596AEBE1-55E7-4246-9ACB-848E85DC29B4%40gladstone.ucsf.edu.
Alex Pico
Dec 1, 2020, 1:47:07 PM12/1/20
to cytoscape-helpdesk
Can you share your CYS file or steps to reproduce the error? The tool works on sample networks, so it is difficult to troubleshoot without seeing the error.
- Alex
To view this discussion on the web visit https://groups.google.com/d/msgid/cytoscape-helpdesk/CAGvYshqqun8VUvYvYXfR8vU2nPF9w2qwaOg6aw6ZBNFirdYcRw%40mail.gmail.com.
Alfred Hargrave
Jan 1, 2021, 9:01:42 AM1/1/21
to cytoscape...@googlegroups.com
Hi Alex,
I've attached an image of my steps. Where can I find the CYS file? Is it cytoscape3.props?
Many thanks, Alfred
To view this discussion on the web visit https://groups.google.com/d/msgid/cytoscape-helpdesk/ED156530-CA7D-4708-8AE0-96ED354D2581%40gladstone.ucsf.edu.
Alfred Hargrave
Jan 1, 2021, 9:04:27 AM1/1/21
to cytoscape...@googlegroups.com
I have just tried it without clicking Analyze as Directed Graph and this has produced statistics and 2 buttons on the right hand side.
If I click "Node degree distirbution", there is a blank error message.
To view this discussion on the web visit https://groups.google.com/d/msgid/cytoscape-helpdesk/ED156530-CA7D-4708-8AE0-96ED354D2581%40gladstone.ucsf.edu.
Alex Pico
Jan 4, 2021, 5:16:36 PM1/4/21
to cytoscape...@googlegroups.com
Great. The steps help. However, if I run those steps on any standard sample network, then it works fine. So, I suspect there is something unique about your network that is exposing a bug in the Analyze Network function. It will be essential to work with your network in order to reproduce and track down the bug.
The CYS file is simply the CYtoscape Session file that is saved to your local hard drive when you when click on the save icon or File > Save. The saved file will be given the .cys extension. Can you email that to me at alex...@gladstone.ucsf.edu?
- Alex
To view this discussion on the web visit https://groups.google.com/d/msgid/cytoscape-helpdesk/CAGvYsho3PDt5K1Q8HJgOKgzQnBaG%3DYT7HZSe4wWhGp%3D_Y%3D%3DZXw%40mail.gmail.com.
Alfred Hargrave
Jan 5, 2021, 5:24:14 AM1/5/21
to cytoscape-helpdesk
Thank you I will send it to you.
Alex Pico
Jan 5, 2021, 1:46:46 PM1/5/21
to cytoscape-helpdesk
Thanks for sharing your session file. A few observations that might be helpful:
1. The network is a protein-protein interaction network (from STRING), so it should not be analyzed as a directed network, i.e., there is no meaning in the "direction" of the connections. For example, RPL19 binds to EIF3K just as much as EIF3K binds to RPL19. So, directed analyses will fail on undirected networks. That examples the initial error... though the app should probably give a more helpful error message! I will file that as a feature request.
2. When running with the option unchecked, it runs just fine (as you also experienced). The two buttons load plots of the distribution of values that you see in the Node Table. This works fine in the latest Cytoscape on my machine. So, this issue does not appear to be specific to your network. The plots rely on CyCharts, which depends upon JavaFX. I suspect you are either running an older version of Cytoscape (get the latest 3.8.2 version here: https://cytoscape.org/download.html) or there is an issus running JavaFX on your machine. Can you confirm the version of Cytoscape? Can you share your OS version? From the screenshots, I'm guessing this might be a linux distro issue and installing JavaFX will resolve it.
3. Not related to your issue, but perhaps still useful, note that anayzer applies to all nodes and edges in the current network view. Your screenshot is zoomed into a corner of the network that you have laid out nicely, but there are a bunch of other nodes and edges below this region. If you want to know the distribution of degree values for just a portion of the network, then drag-select the nodes of interest and create a new subnetwork (File > New Network > From selected nodes, all edges). Then you can run Analyze Network on the subnetwork.
- Alex
Alfred Hargrave
Jan 14, 2021, 12:00:51 PM1/14/21
to cytoscape...@googlegroups.com
Hi Alex, thanks for your reply. Unfortunately I haven't managed to get any of those solutions to work.
My Cytoscape version was 3.8.1, now upgraded to 3.8.2.
The OS I use is Windows 10 Home. I'm not sure how to get JavaFX to install on my computer -- it seems to be something Java 8 uses?
Is there a way to find the node distribution on the STRING website? I can see node count and other statistics under "analysis > Network status" but not node distribution.
Many thanks,
Alfred
To view this discussion on the web visit https://groups.google.com/d/msgid/cytoscape-helpdesk/410769b8-41f9-415f-ac08-74f55caf5b48n%40googlegroups.com.
Alex Pico
Jan 14, 2021, 6:07:55 PM1/14/21
to cytoscape...@googlegroups.com, Cytostaff
Does anyone on this list have experience or suggestions for checking/installing JavaFX on Windows 10 for Cytoscape (e.g., CyCharts)?
- Alex
To view this discussion on the web visit https://groups.google.com/d/msgid/cytoscape-helpdesk/CAGvYshr4yW4avKRp9cKKQBkP74do0nyXq%3DFofG_w02ri-2ONVg%40mail.gmail.com.
Alfred Hargrave
Jan 15, 2021, 5:52:25 AM1/15/21
to cytoscape...@googlegroups.com
I also have Ubuntu by dual boot, so if nothing else works, I could just copy the files over and try doing the node distribution on Ubuntu.
Alfred
To view this discussion on the web visit https://groups.google.com/d/msgid/cytoscape-helpdesk/5996E2EE-F3E8-4D06-B1A1-05AC43D07F13%40gladstone.ucsf.edu.
Scooter Morris
Jan 21, 2021, 11:26:12 AM1/21/21
to cytoscape-helpdesk
Hi Alfred,
What version of Java are you running? By default, we include the modules you need to use JavaFX, but (I believe) we include openJFX modules. You'll need to be running Java 11, and we recommend OpenJDK 11. You shouldn't have to install anything to be JavaFX to work.
-- scooter
Alex Pico
Jan 21, 2021, 12:18:42 PM1/21/21
to cytoscape-helpdesk
Alfred. Another test to help isolate the issue would be to attempt to run CyBrowser (Help>Tutorials should open in CyBrowser). If that works, then JavaFX is working fine and the issue is specific to CyPlot. If not, then JavaFX is still the top suspect.
Alfred Hargrave
Jan 28, 2021, 1:15:56 PM1/28/21
to cytoscape...@googlegroups.com
Hi Scooter, I'll install Java 11. DO you think that will solve the Cytoscape problem on its own?
The version I have is 1.8.0_261
Many thanks, Alfred
Many thanks, Alfred
To view this discussion on the web visit https://groups.google.com/d/msgid/cytoscape-helpdesk/fad1b1f7-907d-48b7-82ca-e04589ecd945n%40googlegroups.com.
Scooter Morris
Feb 4, 2021, 11:43:32 AM2/4/21
to cytoscape-helpdesk
Hi Alfred,
Don't know for sure, but it's a good starting point. Cytoscape depends on the module system in Java 11 to import JavaFX, so it would make sense.
-- scooter
Alfred Hargrave
Feb 4, 2021, 12:07:40 PM2/4/21
to cytoscape...@googlegroups.com
Dear All, I've resolved this issue by analyzing the network in Ubuntu. This did not produce the previous errors. Best, Alfred.
To view this discussion on the web visit https://groups.google.com/d/msgid/cytoscape-helpdesk/99c8584c-e04e-46ab-b859-13f8332ad61bn%40googlegroups.com.
Reply all
Reply to author
Forward
0 new messages
