Simulation of a particular STDP function
49 views
Skip to first unread message
henrymiller....@gmail.com
Jul 18, 2020, 12:59:05 PM7/18/20
to Brian
I refer to the above example. I used the below code and got the following outputs:
-------------------------------------------------------------------------------------------------------------------------------------------------------------
-------------------------------------------------------------------------------------------------------------------------------------------------------------
start_scope()
# Parameters
num_inputs = 100
input_rate = 10*Hz
# Parameters from STDP function
taupre = taupost = 20*ms
wmax = 0.01
Apre = 0.01
Apost = -Apre*1.05
# Time constant
tau = 1*ms
# Construct the poisson neurons
P = PoissonGroup(num_inputs, rates=input_rate)
eqs = '''
dv/dt = -v/tau : 1
'''
# Construct the output neuron
G = NeuronGroup(1, eqs, threshold='v>0.04', reset='v=0', method='exact')
# Define the synapses
S = Synapses(P, G,
'''
w: 1
dapre/dt = -apre/taupre : 1 (event-driven)
dapost/dt = -apost/taupost : 1 (event-driven)
''',
on_pre='''
v_post += w
apre += Apre
w = clip(w+apost, 0, wmax)
''',
on_post='''
apost += Apost
w = clip(w+apre, 0, wmax)
''', method='linear')
S.connect()
# Initialise the weights of the synapses
S.w = 0.01
# State monitor for synapses
M = StateMonitor(S,['w', 'apre', 'apost'], record=True)
# State monitor for output neuron
F = StateMonitor(G,'v',record=True)
# Run simulation for 2 second
run(2*second)
--------------------------------------------------------------------------------------------------------------------------------------------------------
Output:
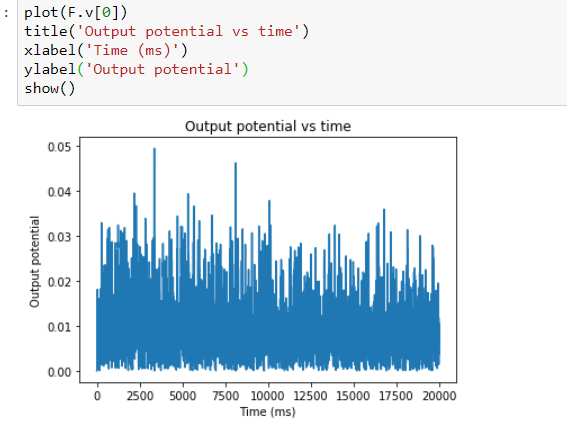
Is this really the correct way to do this? Do the weights really represent the conductances or no? Someone please enlighten me on this. Thanks!
Marcel Stimberg
Aug 4, 2020, 5:33:26 AM8/4/20
to Brian
Hi Henry,
your model looks ok in general, but the weights do not correspond to conductances in the strict sense. You'll find more info on synaptic models in textbooks, but roughly:
- if your synaptic event directly increases the post-synaptic membrane potential, you are using the simplest possible model of a synapse, which is often called "delta synapse" (the post-synaptic membrane potential increases instantaneously)
- If your synaptic event increases a current (and additive term in the equation for v), you have a "current-based synapse" – the increase of the post-synaptic membrane potential is no longer instantaneous, but it is the same independent of the current value of the membrane potential
- If your synaptic event increases a conductance (g in a term like g*(E_syn - v) that is added in the equation for v), you have a "conductance-based synapse" – the increase of the post-synaptic membrane potential now depends on the current value of the membrane potential (bigger effect of the synapse the further away you are from E_syn, the reversal potential of the synapse).
Only in 3) the synaptic weight is a conductance.
Marcel
Reply all
Reply to author
Forward
0 new messages
