Regarding molecular weight analysis in RAW
10 views
Skip to first unread message
Vandanashree Muralidharan
Mar 2, 2022, 12:43:38 AM3/2/22
to BioXTAS RAW
Hi Team,
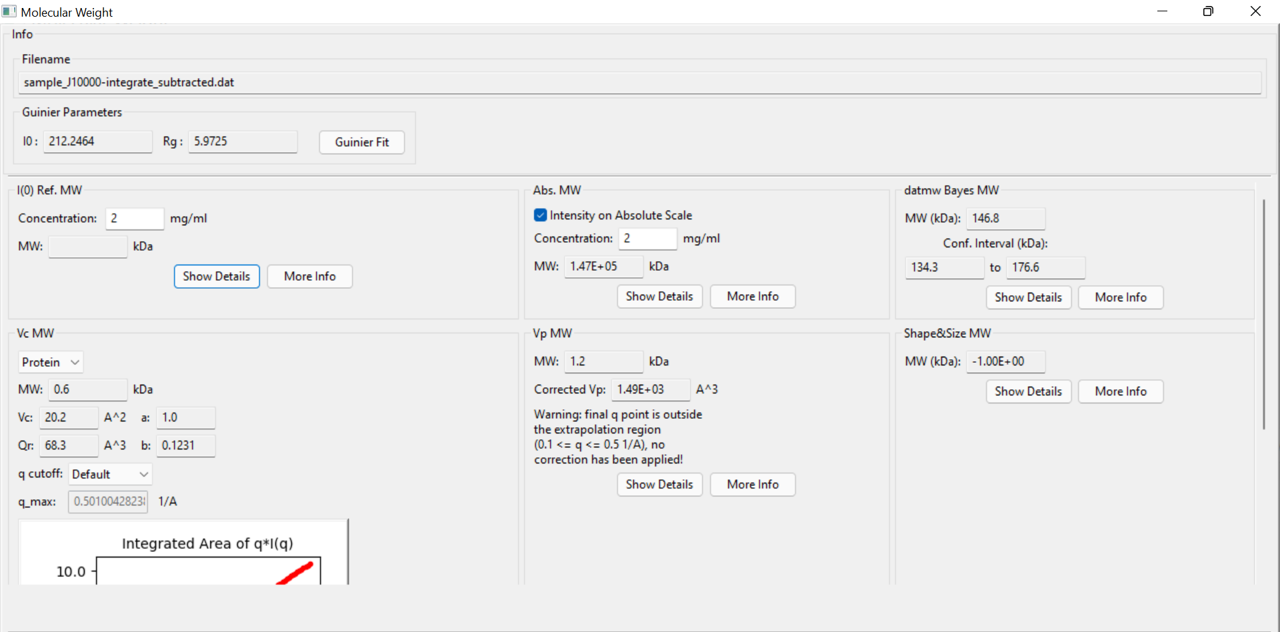
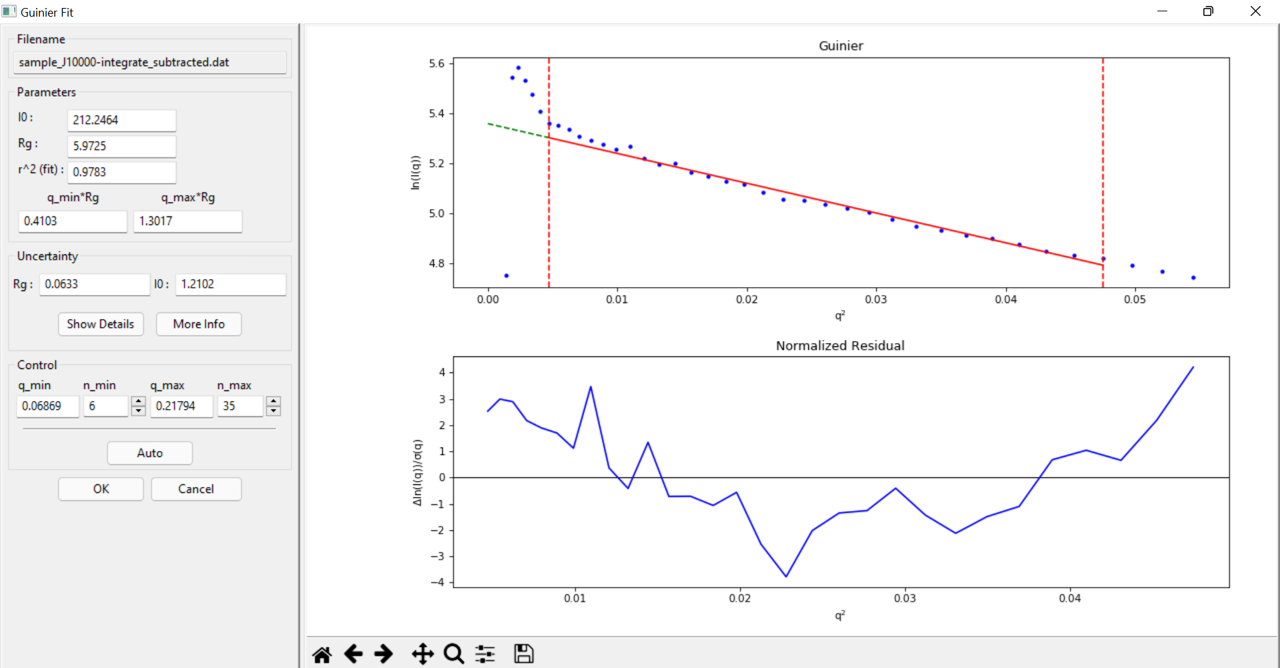
I am Vandana here. While I process the SAXS data for molecular weight estimation, I got certain warning messages, MW based on I(0) Ref MW is not estimated and certain abnormal values in different analysis methods. I have appended here the screenshot of MW panel and Guinier fit for your reference. Kindly suggest how to address this issue. The expected molecular weight is roughly 126 kDa. Thanks.


Jesse Hopkins
Mar 2, 2022, 12:35:38 PM3/2/22
to bioxt...@googlegroups.com
Hi Vandanashree,
It looks like you're using q in 1/nm. Try converting that to 1/Angstrom and see how it all looks. Both the Vc and Vp methods have an inherent assumption of q being in 1/A.
Using either of the concentration dependent methods depends on having a well calibrated system. For the I(0) reference MW you need to have calibrated your system with a known standard, and then set the standard in the RAW configuration. See this tutorial for details:
For the absolute scaling, your data needs to be on an absolute scale, which again depends on the instrument calibration (and, if you're using RAW to reduce your data, as well as analyze it, on having the correct configuration in RAW).
So unless you have those appropriate calibrations for your data the concentration dependent methods of MW calculation may not return useful values.
All the best.
- Jesse
----
Jesse Hopkins, PhD
Deputy Director
BioCAT, Sector 18
Advanced Photon Source
--
You received this message because you are subscribed to the Google Groups "BioXTAS RAW" group.
To unsubscribe from this group and stop receiving emails from it, send an email to bioxtas_raw...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/bioxtas_raw/9e53d251-f97f-48a7-b549-4d244b2a532an%40googlegroups.com.
Reply all
Reply to author
Forward
0 new messages
