ICLV Normalization
221 views
Skip to first unread message
Sally Chen
Mar 20, 2022, 12:21:11 PM3/20/22
to Biogeme
Hi Professor Bierlaire,
I am estimating a hybrid choice model by looking at this example: https://biogeme.epfl.ch/examples/latent/05latentChoiceFull_mc.py
I got some questions regarding the normalization.
- Why are the intercept, beta and sigma of Envir01 all normalized? This is different from what I learned in my class, where I learned to only normalize the factor loading, but no the intercept and sigma.
- I learned that the other way of normalization for LV is by fixing variance term. Is the following correct to do that? sigma_s = Beta('sigma_s', 1, None, None, 1)
Thank you very much for answering the question!
Best,
Sally
Bierlaire Michel
Mar 25, 2022, 9:14:30 AM3/25/22
to sallyc...@gmail.com, Bierlaire Michel, Biogeme
On 19 Mar 2022, at 21:36, Sally Chen <sallyc...@gmail.com> wrote:
Hi Professor Bierlaire,
I am estimating a hybrid choice model by looking at this example: https://biogeme.epfl.ch/examples/latent/05latentChoiceFull_mc.py
I got some questions regarding the normalization.
- Why are the intercept, beta and sigma of Envir01 all normalized?
You need to set the units of the latent variable. Here, we use the first indicator to set the units.
I believe that normalizing the scale should do it as well. This is what we do with utility anyway.
I’ve never tried it though.
- This is different from what I learned in my class, where I learned to only normalize the factor loading, but no the intercept and sigma.
- I learned that the other way of normalization for LV is by fixing variance term. Is the following correct to do that? sigma_s = Beta('sigma_s', 1, None, None, 1)
Thank you very much for answering the question!
Best,Sally
--
You received this message because you are subscribed to the Google Groups "Biogeme" group.
To unsubscribe from this group and stop receiving emails from it, send an email to biogeme+u...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/biogeme/0d102de4-fb1b-4d01-90e6-b64d74a5d414n%40googlegroups.com.
Sally Chen
Apr 13, 2022, 2:16:25 AM4/13/22
to Biogeme
Thanks Prof. Bierlaire!
I have another question about convergence. I'm seeing results with std error of 1.8e+308, even though the robust standard error is computed. However, the cause of termination is saying:
- Cause of termination: Relative gradient = 2.5e-06 <= 6.1e-06
Does this mean the model hasn't converged yet? What are the ways that I can do to make it converge if this is an issue?
Thank you very much!
Bierlaire Michel
Apr 14, 2022, 3:43:29 AM4/14/22
to sallyc...@gmail.com, Bierlaire Michel, Biogeme
On 12 Apr 2022, at 21:43, Sally Chen <sallyc...@gmail.com> wrote:
Thanks Prof. Bierlaire!
I have another question about convergence. I'm seeing results with std error of 1.8e+308, even though the robust standard error is computed. However, the cause of termination is saying:
- Cause of termination: Relative gradient = 2.5e-06 <= 6.1e-06
Does this mean the model hasn't converged yet? What are the ways that I can do to make it converge if this is an issue?
No, it has converged. But there may be numerical issues. In order to calculate the standard errors, Biogeme takes the square root of the diagonal elements of the variance-covariance matrix. If it happens to be negative, the value “1.8e+308” is generated
as a result.
In that case, I always investigate if there is a possible over specification, which typically generates this issue.
However, the model is valid. It is also a good idea to double-check the robust standard errors using bootstrapping.
Thank you very much!
On Friday, March 25, 2022 at 9:14:30 AM UTC-4 michel.b...@epfl.ch wrote:
On 19 Mar 2022, at 21:36, Sally Chen <sallyc...@gmail.com> wrote:
Hi Professor Bierlaire,
I am estimating a hybrid choice model by looking at this example: https://biogeme.epfl.ch/examples/latent/05latentChoiceFull_mc.py
I got some questions regarding the normalization.
- Why are the intercept, beta and sigma of Envir01 all normalized?
You need to set the units of the latent variable. Here, we use the first indicator to set the units.I believe that normalizing the scale should do it as well. This is what we do with utility anyway.I’ve never tried it though.
- This is different from what I learned in my class, where I learned to only normalize the factor loading, but no the intercept and sigma.
- I learned that the other way of normalization for LV is by fixing variance term. Is the following correct to do that? sigma_s = Beta('sigma_s', 1, None, None, 1)
Thank you very much for answering the question!
Best,Sally
--
You received this message because you are subscribed to the Google Groups "Biogeme" group.
To unsubscribe from this group and stop receiving emails from it, send an email to biogeme+u...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/biogeme/0d102de4-fb1b-4d01-90e6-b64d74a5d414n%40googlegroups.com.
--
You received this message because you are subscribed to the Google Groups "Biogeme" group.
To unsubscribe from this group and stop receiving emails from it, send an email to biogeme+u...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/biogeme/0db93f4f-e1f9-4c23-abec-69cae8351177n%40googlegroups.com.
Sally Chen
Apr 20, 2022, 12:04:50 PM4/20/22
to Biogeme
Hi prof. Bierlaire,
Thank you for the answer! One more question: how is the robust standard error computed? Is the robust SE valid?
Thanks!
Best,
Sally
Bierlaire Michel
Apr 20, 2022, 12:06:49 PM4/20/22
to sallyc...@gmail.com, Bierlaire Michel, Biogeme
On 20 Apr 2022, at 15:39, Sally Chen <sallyc...@gmail.com> wrote:
Hi prof. Bierlaire,
Thank you for the answer! One more question: how is the robust standard error computed?
See Appendix B of this document: https://transp-or.epfl.ch/documents/technicalReports/Bier20.pdf
Is the robust SE valid?
It should, in principle.
To view this discussion on the web visit https://groups.google.com/d/msgid/biogeme/2b67b1cb-a54f-4280-bdf2-73644e70c00an%40googlegroups.com.
Sally Chen
Oct 3, 2022, 2:11:48 AM10/3/22
to Biogeme
Hi Prof. Bierlaire,
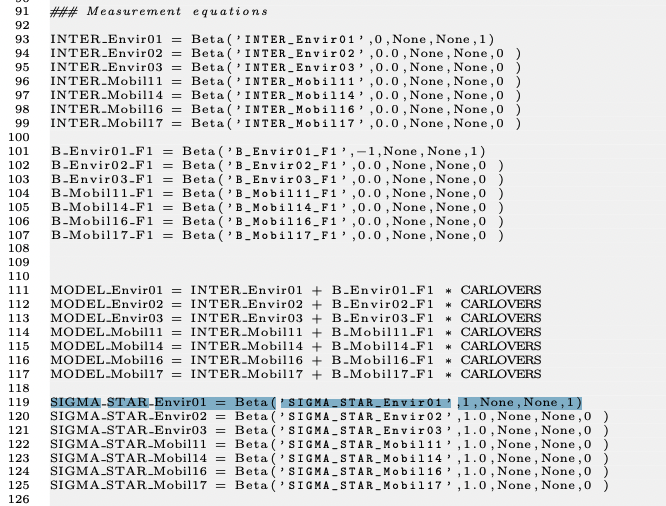
Sorry that I'm reopening this thread. I just read the section about measurement equation (ordered probit) in this document: https://transp-or.epfl.ch/documents/technicalReports/Bier18b.pdf. I have three additional questions:
- I understand the necessity to set B_Envir01_F1 to be fixed, but why INTER_Envir01?
- In this model, we have additional latent response variables (e.g. Envir01*). Since we need to fix the scale of every latent variable, shouldn't their scale also be fixed? Meaning that shouldn't all of the sigma (line 119 - 125) be fixed to 1? SIGMA_STAR_Envir01 is fixed to 1, but why not the others?
- Why in the structural equation, the sigma of car lover LV is not estimated? If we fix the loading of one of the indicators to -1/1, then that is estimable right?
Thank you very much for answering these questions!
Best,
Sally

On Friday, March 25, 2022 at 9:14:30 AM UTC-4 michel.b...@epfl.ch wrote:
Bierlaire Michel
Oct 3, 2022, 2:18:18 AM10/3/22
to sallyc...@gmail.com, Bierlaire Michel, Biogeme
On 3 Oct 2022, at 05:24, Sally Chen <sallyc...@gmail.com> wrote:
Hi Prof. Bierlaire,
Sorry that I'm reopening this thread. I just read the section about measurement equation (ordered probit) in this document: https://transp-or.epfl.ch/documents/technicalReports/Bier18b.pdf. I have three additional questions:
- I understand the necessity to set B_Envir01_F1 to be fixed, but why INTER_Envir01?
The first sets the scale, the second sets the location.
- In this model, we have additional latent response variables (e.g. Envir01*). Since we need to fix the scale of every latent variable, shouldn't their scale also be fixed? Meaning that shouldn't all of the sigma (line 119 - 125) be fixed to 1? SIGMA_STAR_Envir01 is fixed to 1, but why not the others?
Because their variances are not necessarily the same.
- Why in the structural equation, the sigma of car lover LV is not estimated? If we fix the loading of one of the indicators to -1/1, then that is estimable right?
It is estimated. See http://biogeme.epfl.ch/examples/latent/05latentChoiceFull.py
Thank you very much for answering these questions!
Best,Sally
<Screen Shot 2022-10-02 at 23.18.56.png>
On Friday, March 25, 2022 at 9:14:30 AM UTC-4 michel.b...@epfl.ch wrote:
On 19 Mar 2022, at 21:36, Sally Chen <sallyc...@gmail.com> wrote:
Hi Professor Bierlaire,
I am estimating a hybrid choice model by looking at this example: https://biogeme.epfl.ch/examples/latent/05latentChoiceFull_mc.py
I got some questions regarding the normalization.
- Why are the intercept, beta and sigma of Envir01 all normalized?
You need to set the units of the latent variable. Here, we use the first indicator to set the units.I believe that normalizing the scale should do it as well. This is what we do with utility anyway.I’ve never tried it though.
- This is different from what I learned in my class, where I learned to only normalize the factor loading, but no the intercept and sigma.
- I learned that the other way of normalization for LV is by fixing variance term. Is the following correct to do that? sigma_s = Beta('sigma_s', 1, None, None, 1)
Thank you very much for answering the question!
Best,Sally
--
You received this message because you are subscribed to the Google Groups "Biogeme" group.
To unsubscribe from this group and stop receiving emails from it, send an email to biogeme+u...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/biogeme/0d102de4-fb1b-4d01-90e6-b64d74a5d414n%40googlegroups.com.
--
You received this message because you are subscribed to the Google Groups "Biogeme" group.
To unsubscribe from this group and stop receiving emails from it, send an email to biogeme+u...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/biogeme/6e296553-d0b9-4c1d-865e-81f77edbd5b7n%40googlegroups.com.
Reply all
Reply to author
Forward
Message has been deleted
Message has been deleted
0 new messages
