Substitution model selection in BEAST
Ching Ching WEE
The best-fit model according to AICc using ModelFinder is : HIVw+F+I+G4
However, may I know which substitution model should I choose as both BEAST version did not have HIVw option as shown in the printscreen below?
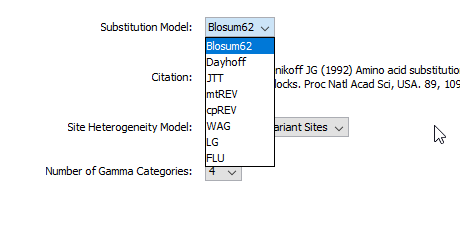
BEASTv.1.10.4
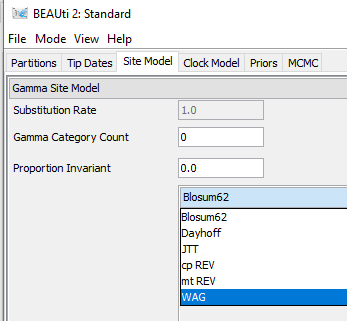
BEASTv2.6.7
In addition, how to specify 'F' (empirical frequencies)? I only see empirical frequencies option if the HKY substitution model is chosen (nucleotide). However, my data type is amino acids.
Remco Bouckaert

On 3/05/2022, at 11:43 AM, Ching Ching WEE <wching...@gmail.com> wrote:
Dear BEAST User
The best-fit model according to AICc using ModelFinder is : HIVw+F+I+G4
However, may I know which substitution model should I choose as both BEAST version did not have HIVw option as shown in the printscreen below?
<2022-05-03 02_35_05-BEAUtiv.1.10.4.png>
BEASTv.1.10.4
<2022-05-03 02_35_47-BEAUti 2_ Standard.png>
BEASTv2.6.7
In addition, how to specify 'F' (empirical frequencies)? I only see empirical frequencies option if the HKY substitution model is chosen (nucleotide). However, my data type is amino acids.
--
You received this message because you are subscribed to the Google Groups "beast-users" group.
To unsubscribe from this group and stop receiving emails from it, send an email to beast-users...@googlegroups.com.
To view this discussion on the web visit https://groups.google.com/d/msgid/beast-users/4a737e73-ed09-4a13-aaa3-05e643d04d48n%40googlegroups.com.
<2022-05-03 02_35_05-BEAUtiv.1.10.4.png><2022-05-03 02_35_47-BEAUti 2_ Standard.png>
