setting priors for Path Sampling/Stepping-stone sampling
98 views
Skip to first unread message
Samuel Zimmerman
Jan 2, 2022, 2:03:24 PM1/2/22
to beast-users
Hi all,
I recently ran Beast to perform divergence dating on a species of bacteria using several different tree priors (coalescent constant population, coalescent exponential population, coalescent bayesian skyline, and coalescent exponential skyline).
I would like to compare the marginal likelihoods of these models to each other using path sampling, but realized that the default priors for these models need to be changed when doing model selection because of improper priors. I understand that you need to change the popSize and ePopSize parameters for the constant and exponential population models respectively.
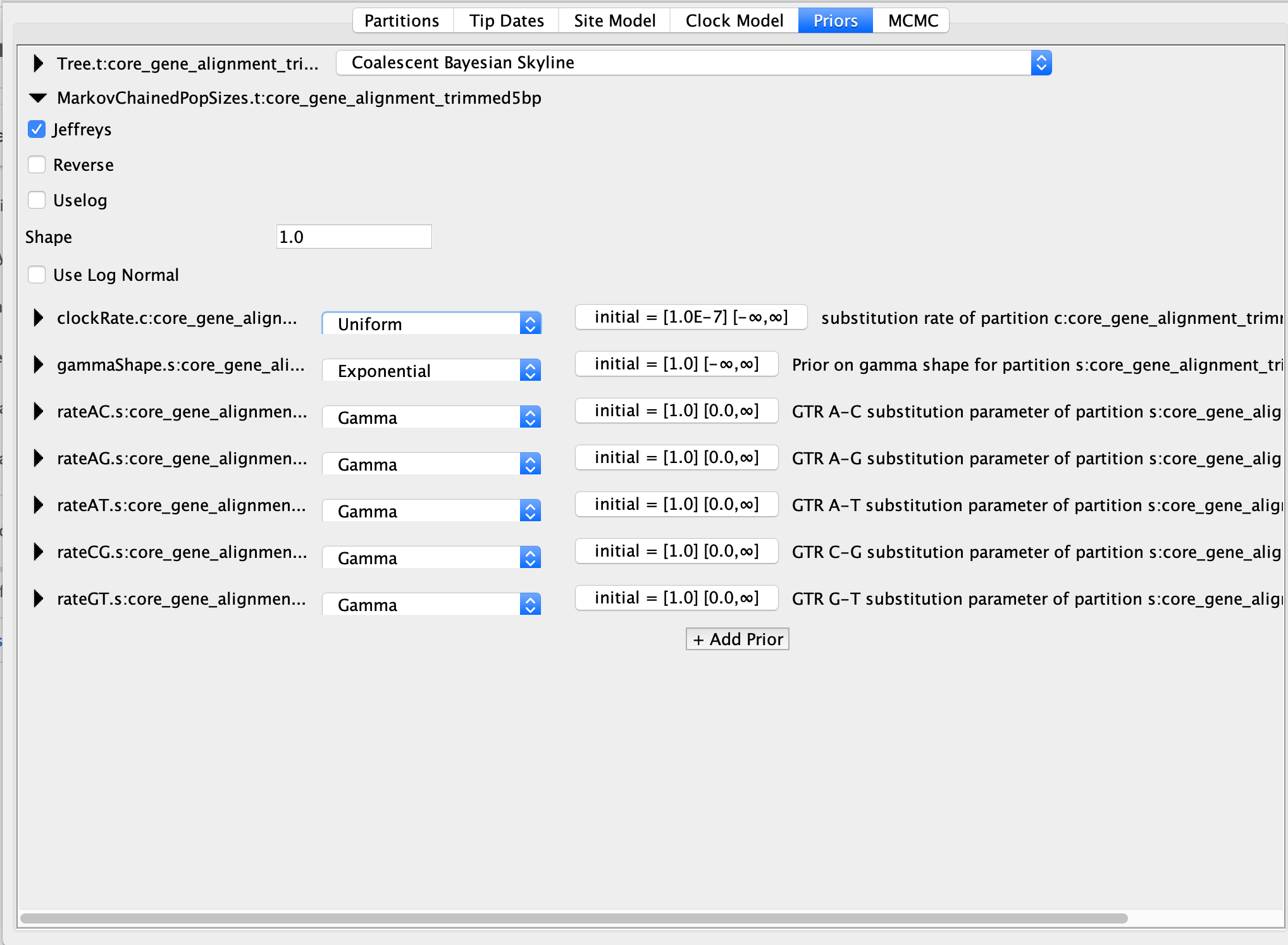
However, for the Coalescent Bayesian Skyline model, there is a parameter called "MarkovChainedPopSizes" that is set to be a Jeffreys prior by default. From my understanding Jeffreys is an improper prior but I am not sure. Should I set this to "Uselog" instead? Attached you can find a screen shot illustrating the options available.
Thank you very much!
Best,
Sam
Sam

Reply all
Reply to author
Forward
0 new messages
