How to set conditions with $ such that featureData() has separate columns for the conditions
68 views
Skip to first unread message
Dmitry Leontyev
Feb 11, 2022, 1:22:15 PM2/11/22
to Cardinal MSI Help
Hello everyone,

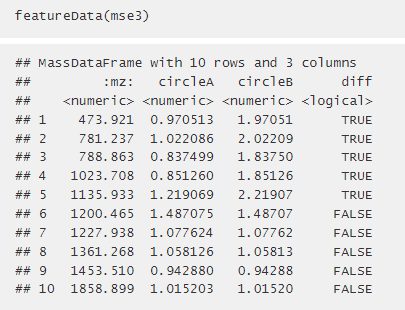
I am following the class comparison examples (http://bioconductor.org/packages/release/bioc/vignettes/Cardinal/inst/doc/Cardinal-2-stats.html#class-comparison) and attempting to adapt it to my data.


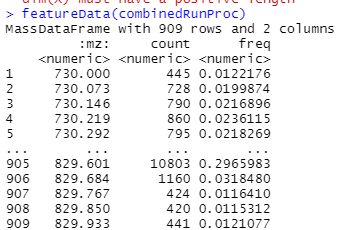
In the examples above, when makeFactor is called, the $ operator is used to split conditions in a manner which I cannot figure out how to adapt it to my code. When featureData() is called on mse3, there are separate columns for "circleA" and "circleB". When I run it on my data set, there is just one column (see below). I want to split my columns like was done in the example, but am having trouble figuring out how to do so. I have included my code below. If anyone could point out how I could accomplish this I would greatly appreciate it.
Thank you,
Dmitry

--------MY CODE---------
library(Cardinal)
#defines paths
pathtest1 <- "C:/Users/Dima/Downloads/DANdimer.imzML"
pathtest2 <- "C:/Users/Dima/Downloads/LPC160.imzML"
#loads data
test1 <- readMSIData(pathtest1, mass.range=c(730,830), resolution=100, units="ppm", attach.only=TRUE)
test2 <- readMSIData(pathtest2, mass.range=c(730,830), resolution=100, units="ppm", attach.only=TRUE)
#sets centroided as TRUE, which is necessary for combination and combines runs
centroided(test1) <- TRUE
centroided(test2) <- TRUE
combinedRun <- Cardinal::combine(test1,test2)
#pre-process (in future might want to select ROIs before pre-process to reduce processing time)
combinedRunPre <- combinedRun %>% normalize(method="rms", delay=TRUE) %>% peakFilter(freq.min=0.01)
processingData(combinedRunPre)
combinedRunProc <- process(combinedRunPre,centroided=TRUE,plot=TRUE)
#ROI selection and subsetting and combining into regions
sampleA <- selectROI(combinedRunProc, mz=734.569, subset=run(combinedRunProc) == "DANdimer")
sampleB <- selectROI(combinedRunProc, mz=734.569, subset=run(combinedRunProc) == "LPC160")
regions <- makeFactor(DANdimer=sampleA, LPC160=sampleB)
#defines paths
pathtest1 <- "C:/Users/Dima/Downloads/DANdimer.imzML"
pathtest2 <- "C:/Users/Dima/Downloads/LPC160.imzML"
#loads data
test1 <- readMSIData(pathtest1, mass.range=c(730,830), resolution=100, units="ppm", attach.only=TRUE)
test2 <- readMSIData(pathtest2, mass.range=c(730,830), resolution=100, units="ppm", attach.only=TRUE)
#sets centroided as TRUE, which is necessary for combination and combines runs
centroided(test1) <- TRUE
centroided(test2) <- TRUE
combinedRun <- Cardinal::combine(test1,test2)
#pre-process (in future might want to select ROIs before pre-process to reduce processing time)
combinedRunPre <- combinedRun %>% normalize(method="rms", delay=TRUE) %>% peakFilter(freq.min=0.01)
processingData(combinedRunPre)
combinedRunProc <- process(combinedRunPre,centroided=TRUE,plot=TRUE)
#ROI selection and subsetting and combining into regions
sampleA <- selectROI(combinedRunProc, mz=734.569, subset=run(combinedRunProc) == "DANdimer")
sampleB <- selectROI(combinedRunProc, mz=734.569, subset=run(combinedRunProc) == "LPC160")
regions <- makeFactor(DANdimer=sampleA, LPC160=sampleB)
--------------------------------
Reply all
Reply to author
Forward
0 new messages
