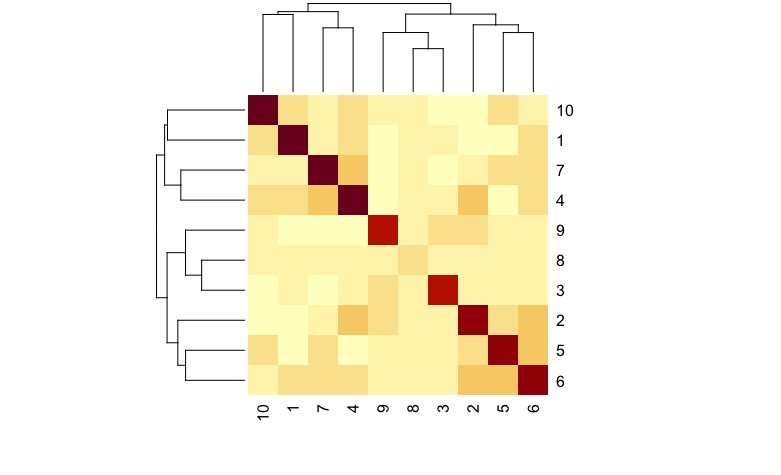
kinship matrix plot
Atoosa Samani
Karl Broman
Dan Gatti
You might look at the degree of homozygosity in each sample. I suspect that sample 8 has lower homozygosity than the other samples.
I was confused by this same thing for a long time. When allele sharing within the same sample is estimated, you compare allele1 with allele2 at one locus. If they’re both ‘A’, then the locus shares an allele. If allele1 is ‘A’ and allele2 is ‘C’, then they don’t share an allele at that locus. So if you have lots of homozygous loci, your self-kinship will be closer to 1. As the number of heterozygous loci increases, you’ll see lower self-kinship numbers.
Does that help?
Dan
--
You received this message because you are subscribed to the Google Groups "R/qtl2 discussion" group.
To unsubscribe from this group and stop receiving emails from it, send an email to
rqtl2-disc+...@googlegroups.com.
To view this discussion on the web visit
https://groups.google.com/d/msgid/rqtl2-disc/5b5b8c3b-701d-43f1-91f3-8ee414853f1fn%40googlegroups.com.
The information in this email, including attachments, may be confidential and is intended solely for the addressee(s). If you believe you received this email by mistake, please notify the sender by return email as soon as possible.
