Chickenosaurus
22 views
Skip to first unread message
John F Sowa
May 4, 2016, 10:32:51 AM5/4/16
to ontolog-forum
Biological types were the inspiration for "natural kinds"
that are organized in well-defined trees with no cross links.
But artifacts are much harder to classify because people keep
inventing new versions and combinations. Geological formations
are also harder to classify because the genetic constraints
that keep species distinct don't apply to geology.
Gene editing is now blurring the distinction between natural
kinds and artifacts. Genes can even be transplanted between
plants and animals.
Today, some scientists are even "reverse engineering" evolution.
Since birds evolved from dinosaurs, they're experimenting with
the easily available chicken eggs to create a "chickenosaurus":
http://motherboard.vice.com/read/why-scientists-on-two-continents-are-growing-dinosaur-parts-on-chickens-McGill-University-Hans-Larsson?trk_source=recommended
This is just one of many reasons why trees are not a good way
to organize ontological categories.
John
that are organized in well-defined trees with no cross links.
But artifacts are much harder to classify because people keep
inventing new versions and combinations. Geological formations
are also harder to classify because the genetic constraints
that keep species distinct don't apply to geology.
Gene editing is now blurring the distinction between natural
kinds and artifacts. Genes can even be transplanted between
plants and animals.
Today, some scientists are even "reverse engineering" evolution.
Since birds evolved from dinosaurs, they're experimenting with
the easily available chicken eggs to create a "chickenosaurus":
http://motherboard.vice.com/read/why-scientists-on-two-continents-are-growing-dinosaur-parts-on-chickens-McGill-University-Hans-Larsson?trk_source=recommended
This is just one of many reasons why trees are not a good way
to organize ontological categories.
John
Ravi Sharma
May 4, 2016, 11:31:14 AM5/4/16
to ontolog-forum
John
Great examples of changes and hybrid constructs, fusion across new lines and cross products.
Traditionally Taxonomies are integral to botany and I thought ontologies would be helpful if cross linkages were found using them among phylum, genera, species, etc.?
What you mention are often exceptions but we realize that like Genetically Modified food, these would new types would intrude and become more the new "natural" way and thus confuse the erstwhile ok ontologies!.
Regards,
John
--
All contributions to this forum by its members are made under an open content license, open publication license, open source or free software license. Unless otherwise specified, all Ontolog Forum content shall be subject to the Creative Commons CC-BY-SA 4.0 License or its successors.
--- You received this message because you are subscribed to the Google Groups "ontolog-forum" group.
To unsubscribe from this group and stop receiving emails from it, send an email to ontolog-foru...@googlegroups.com.
To post to this group, send email to ontolo...@googlegroups.com.
Visit this group at https://groups.google.com/group/ontolog-forum.
To view this discussion on the web visit https://groups.google.com/d/msgid/ontolog-forum/ced80c16-1be6-e495-0559-ab520b4048db%40bestweb.net.
For more options, visit https://groups.google.com/d/optout.
--
Chris Mungall
May 4, 2016, 11:52:38 AM5/4/16
to ontolog-forum
In fact even in the absence of human intervention, trees in biology are
a huge simplification. Horizontal gene transfer is a natural phenomenon.
Bacteria do it all the time, and there have been multiple events of
bacterial genes being introduced into complex organisms, including our
own lineage.
This figure shows some of the HGT events early in the plant lineage,
including functions that were transferred across:
http://www.nature.com/ncomms/journal/v3/n10/fig_tab/ncomms2148_F4.html
I don't know anyone in bio-ontologies who still thinks ontologies should
be trees. The standard engineering methodology is: assert a tree, infer
a graph
David Poole
May 4, 2016, 12:48:49 PM5/4/16
to ontolo...@googlegroups.com
Hi All,
John is absolutely right (as he normally is).
We have been having lots of success with what have been called "Aristotelian Definitions", where all classes are all defined in terms of a genus (a superclass) and diffentia (what differentiates this class from other subclasses). This typically does not lead to a tree, but to a lattice-like structure (it isn't a lattice as as not all properties are defined for all classes). This is a natural way to define many ontologies.
We are using such ontologies in probabilistic reasoning; conditioning posterior distributions on observations where the terms in the queries and observations are all defined in ontologies. Differentia lead to random variables.
We were inspired by this great quote of Aristotle:
If genera are different and co-ordinate, their differentiae are themselves
different in kind. Take as an instance the genus ``animal'' and the
genus ``knowledge''. ``With feet'', ``two-footed'', ``winged'', ``aquatic'',
are differentiae of ``animal''; the species of knowledge are not distinguished
by the same differentiae. One species of knowledge does not differ
from another in being ``two-footed''.
Aristotle (350 B.C.). Categories. Translated by E. M. Edghill, http://www.classicallibrary.org/aristotle/categories/index.htm
See: David Poole, Clinton Smyth and Rita Sharma, Ontology Design for Scientific Theories That Make Probabilistic Predictions, IEEE Intelligent Systems, Special Issue on Semantic Scientific Knowledge Integration - Jan/Feb 2009, pages 27-36 http://www.cs.ubc.ca/~poole/papers/PooleSmythSharma2009.pdf
There is other follow-up work on my web site.
David
David Poole,
Department of Computer Science,
University of British Columbia,
http://cs.ubc.ca/~poole
po...@cs.ubc.ca
John is absolutely right (as he normally is).
We have been having lots of success with what have been called "Aristotelian Definitions", where all classes are all defined in terms of a genus (a superclass) and diffentia (what differentiates this class from other subclasses). This typically does not lead to a tree, but to a lattice-like structure (it isn't a lattice as as not all properties are defined for all classes). This is a natural way to define many ontologies.
We are using such ontologies in probabilistic reasoning; conditioning posterior distributions on observations where the terms in the queries and observations are all defined in ontologies. Differentia lead to random variables.
We were inspired by this great quote of Aristotle:
If genera are different and co-ordinate, their differentiae are themselves
different in kind. Take as an instance the genus ``animal'' and the
genus ``knowledge''. ``With feet'', ``two-footed'', ``winged'', ``aquatic'',
are differentiae of ``animal''; the species of knowledge are not distinguished
by the same differentiae. One species of knowledge does not differ
from another in being ``two-footed''.
Aristotle (350 B.C.). Categories. Translated by E. M. Edghill, http://www.classicallibrary.org/aristotle/categories/index.htm
See: David Poole, Clinton Smyth and Rita Sharma, Ontology Design for Scientific Theories That Make Probabilistic Predictions, IEEE Intelligent Systems, Special Issue on Semantic Scientific Knowledge Integration - Jan/Feb 2009, pages 27-36 http://www.cs.ubc.ca/~poole/papers/PooleSmythSharma2009.pdf
There is other follow-up work on my web site.
David
Department of Computer Science,
University of British Columbia,
http://cs.ubc.ca/~poole
po...@cs.ubc.ca
John F Sowa
May 6, 2016, 1:50:01 AM5/6/16
to ontolo...@googlegroups.com, upr...@gmail.com
Ravi, Chris, and David,
Basic point: a single-inheritance tree is not adequate to
represent the open-ended variety of ways of classifying anything.
RS
the single-inheritance trees used in biology. But that is
only one way of classifying living things.
Note that the words 'tree' and 'bush' describe the physical forms.
But the same plant can be trimmed as a bush or allowed to grow
into a tree.
Many other classifications are orthogonal to the genetic trees:
evergreen vs deciduous, annual vs perennial, crop vs weed, etc.
In general, any plant that grows where you don't want it can be
called a weed. A dandelion can be a weed or a salad green.
For example, the single family called Rosaceae (the rose family)
has over 2,000 species. Some are evergreen, deciduous, annuals,
perennials, trees, shrubs, herbs, delicious fruits, ornamental
flowers, crops, or noxious weeds.
Examples: roses, apples, pears, cherries, plums, peaches,
raspberries, strawberries, almonds, hawthorns...
Note that farmers, who work with plants and animals every day, normally
use the more specialized classifications. They're more interested in
the crops, weeds, and pests they encounter every day than the complete
classification of every living thing.
CM
> This figure shows some of the HGT (Horizontal Gene Transfers) early in
article. But note that most of those transfers are from bacteria to
more complex life forms. None of those possibilities were imagined
until late in the 20th century.
At the higher levels of plants and animals, trees are still
the primary way of classifying inheritance. But every domain
of interest requires multiple classifications of living things
that cut across the trees in multiple ways for different purposes.
DP
Yes. You need multiple-inheritance, which leads to a general
partial order.
DP
are undefined for many of the classes. The algorithms for
Formal Concept Analysis (FCA), will always generate a lattice.
See the FCA homepage for more info, open source software, etc:
http://www.fcahome.org.uk/fca.html
In fact, it's fun to play around with the FCA lattices generated
from WordNet and Roget's Thesaurus. You can pick any word and
get a little lattice for the WordNet neighborhood or the Roget
neighborhood around that word. Go to either or both of
http://www.ketlab.org.uk/wordnet.html
http://www.ketlab.org.uk/roget.html
Pick any English word and compare the lattices generated for
WordNet and Roget's Thesaurus.
You could download the software and generate similar lattices
for the terms in your ontologies. If you have any questions,
contact Uta Priss (on cc list above).
John
Basic point: a single-inheritance tree is not adequate to
represent the open-ended variety of ways of classifying anything.
RS
> I thought ontologies would be helpful if cross linkages were
> found using them among phylum, genera, species, etc.?
Names, such as phylum, etc., are useful for the levels of
> found using them among phylum, genera, species, etc.?
the single-inheritance trees used in biology. But that is
only one way of classifying living things.
Note that the words 'tree' and 'bush' describe the physical forms.
But the same plant can be trimmed as a bush or allowed to grow
into a tree.
Many other classifications are orthogonal to the genetic trees:
evergreen vs deciduous, annual vs perennial, crop vs weed, etc.
In general, any plant that grows where you don't want it can be
called a weed. A dandelion can be a weed or a salad green.
For example, the single family called Rosaceae (the rose family)
has over 2,000 species. Some are evergreen, deciduous, annuals,
perennials, trees, shrubs, herbs, delicious fruits, ornamental
flowers, crops, or noxious weeds.
Examples: roses, apples, pears, cherries, plums, peaches,
raspberries, strawberries, almonds, hawthorns...
Note that farmers, who work with plants and animals every day, normally
use the more specialized classifications. They're more interested in
the crops, weeds, and pests they encounter every day than the complete
classification of every living thing.
CM
> This figure shows some of the HGT (Horizontal Gene Transfers) early in
> the plant lineage, including functions that were transferred across:
> http://www.nature.com/ncomms/journal/v3/n10/fig_tab/ncomms2148_F4.html
Thanks for the reference. That's a good diagram and an interesting
> http://www.nature.com/ncomms/journal/v3/n10/fig_tab/ncomms2148_F4.html
article. But note that most of those transfers are from bacteria to
more complex life forms. None of those possibilities were imagined
until late in the 20th century.
At the higher levels of plants and animals, trees are still
the primary way of classifying inheritance. But every domain
of interest requires multiple classifications of living things
that cut across the trees in multiple ways for different purposes.
DP
> We have been having lots of success with what have been called
> "Aristotelian Definitions", where all classes are all defined in
> terms of a genus (a superclass) and diffentia (what differentiates
> this class from other subclasses). This typically does not lead
> to a tree, but to a lattice-like structure...
> "Aristotelian Definitions", where all classes are all defined in
> terms of a genus (a superclass) and diffentia (what differentiates
> this class from other subclasses). This typically does not lead
Yes. You need multiple-inheritance, which leads to a general
partial order.
DP
> (it isn't a lattice as as not all properties are defined for
> all classes). This is a natural way to define many ontologies.
Actually, you can still get a lattice, even if many properties
> all classes). This is a natural way to define many ontologies.
are undefined for many of the classes. The algorithms for
Formal Concept Analysis (FCA), will always generate a lattice.
See the FCA homepage for more info, open source software, etc:
http://www.fcahome.org.uk/fca.html
In fact, it's fun to play around with the FCA lattices generated
from WordNet and Roget's Thesaurus. You can pick any word and
get a little lattice for the WordNet neighborhood or the Roget
neighborhood around that word. Go to either or both of
http://www.ketlab.org.uk/wordnet.html
http://www.ketlab.org.uk/roget.html
Pick any English word and compare the lattices generated for
WordNet and Roget's Thesaurus.
You could download the software and generate similar lattices
for the terms in your ontologies. If you have any questions,
contact Uta Priss (on cc list above).
John
John Bottoms
May 12, 2016, 1:36:53 PM5/12/16
to ontolo...@googlegroups.com
All,
In a discussion a friend of mine, he told me that there is a trend in China to create strictly visual technical manuals. I was already familiar with the extensive photographs documenting the assembly of the Tiny Boy 3D printer from Singapore. It is assembled by young students and there are about 250 photos in the sequence. In true Chinese style the pictures are view right to left. That started me looking for examples of visual manuals in English.
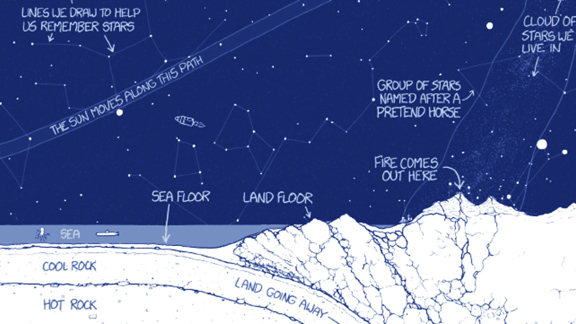
I stumbled across Randall Munroe's book "Thing Explainer". He is the creator of the xkcd comic strip. The premise of his book is to illustrate concepts using just words from the (English, CNL) most common 1000 word list. This interested me because it serves as a way to visually extend a vocabulary and create a technical vocabulary.
The word list URL is:
An image from the book "Thing Explainer" is below.
In a discussion a friend of mine, he told me that there is a trend in China to create strictly visual technical manuals. I was already familiar with the extensive photographs documenting the assembly of the Tiny Boy 3D printer from Singapore. It is assembled by young students and there are about 250 photos in the sequence. In true Chinese style the pictures are view right to left. That started me looking for examples of visual manuals in English.
I stumbled across Randall Munroe's book "Thing Explainer". He is the creator of the xkcd comic strip. The premise of his book is to illustrate concepts using just words from the (English, CNL) most common 1000 word list. This interested me because it serves as a way to visually extend a vocabulary and create a technical vocabulary.
The word list URL is:
http://splasho.com/upgoer5/phpspellcheck/dictionaries/1000.dicin
An image from the book "Thing Explainer" is below.

After thinking about it for a while it occurred
to me that while this is useful to a certain community, it is
not strictly a visual ontology. First, it addresses solely
elements and is more of a lexicon. Attributes are missing.
In looking for a better example I found a poster of musical instruments. The image is large because it represents a complete taxonomy for the well known instruments. I included it in the original size because it is important to see how the hierarchy is represented. Due to its size, I have attached it separately rather than embedding it.
In searching to fill the voice for Processes, I came across Step-by-Step illustrations. These again are not strictly visual but they go a long way toward that goal. It appears that they are used extensively by Ikea to show the assembly of their furniture. Below is an example of a Process illustration borrowed from "Nature".
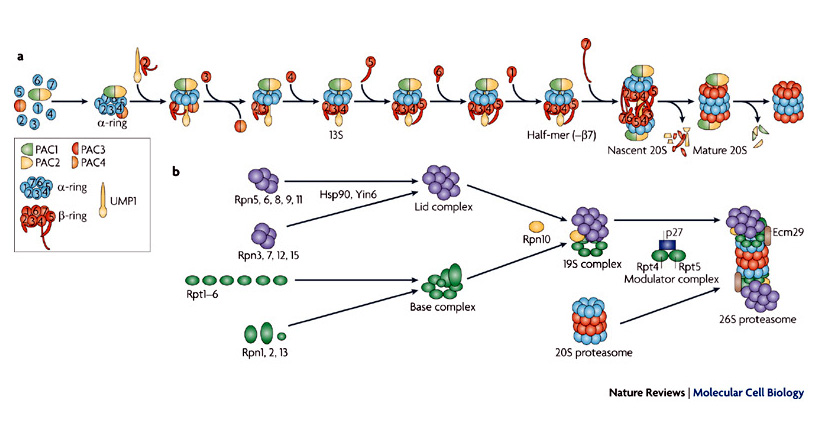
The missing part of all of this is the attributes and
properties. There may be constructs or visual supplements that
can convey the necessary information. I'm still working on that
part.
And finally, since there are three basic ways people learn, Visual, Auditory and Kinesthetic, there are more questions remaining in this study. What questions do you see?
-John Bottoms
Concord, MA USA
In looking for a better example I found a poster of musical instruments. The image is large because it represents a complete taxonomy for the well known instruments. I included it in the original size because it is important to see how the hierarchy is represented. Due to its size, I have attached it separately rather than embedding it.
In searching to fill the voice for Processes, I came across Step-by-Step illustrations. These again are not strictly visual but they go a long way toward that goal. It appears that they are used extensively by Ikea to show the assembly of their furniture. Below is an example of a Process illustration borrowed from "Nature".

And finally, since there are three basic ways people learn, Visual, Auditory and Kinesthetic, there are more questions remaining in this study. What questions do you see?
-John Bottoms
Concord, MA USA
rrov...@buffalo.edu
May 13, 2016, 8:48:30 PM5/13/16
to ontolog-forum, jo...@firststarsystems.com
Good topic, John. We need more discussion on the visual and ontology.
I would go one step further and move from static visual content to video or animation (i.e., dynamic visual content).
Experiences i've had in another discipline may be helpful. In my maritime safety volunteer work, I've created and used visual study guides as early as '06 and late as early this year, an animated example being a simple power point. (Essentially an exercise in instructional design, without even knowing it.) From a pedagogical and self-study standpoint, they are and have been helpful, particularly to people who are visual or kinesthetic (I suspect a connection) learners or who otherwise love the visual element, such as myself.
I would go one step further and move from static visual content to video or animation (i.e., dynamic visual content).
Experiences i've had in another discipline may be helpful. In my maritime safety volunteer work, I've created and used visual study guides as early as '06 and late as early this year, an animated example being a simple power point. (Essentially an exercise in instructional design, without even knowing it.) From a pedagogical and self-study standpoint, they are and have been helpful, particularly to people who are visual or kinesthetic (I suspect a connection) learners or who otherwise love the visual element, such as myself.
1) Question: as you mentioned it, what's the difference between attributes and property? (e.g. property=value of attribute, etc.) Can you give an example for the current visual topic?
2) It's interesting to think how properties/attributes can be visually represented. As it is, we immediately grasp the properties when we see the image. We see the whole--each visual element--and apprehend the properties (shape, color, dimension, size, ...). One way I was able to memorize the specs of some maritime vessels was in mentally memorizing the visual guides I would create: on one page I had the vessel (the whole/totality) along with it's features (the property/attributes), including some relational-features. When wanting to recall (or when being tested!) a particular spec (e.g., freeboard at the bow), I sometimes would mentally recall the whole picture.
3) I believe conveying a processual aspect is critical, particularly for instructional, training, and self-learning. Mentally visualizing and cogitating the whole, the gestalt, the totality of the process is important. I believe this is analogous to hearing a story, to being captured by an story, and dialectic (Plato, not Aristotle, had it right!) Again, during my maritime work and training, I found and find it helpful (often necessary) to be aware of the big-picture of the process, e.g., the goal the reason for the it all, and work through the steps and procedures, in the process.
3) I believe conveying a processual aspect is critical, particularly for instructional, training, and self-learning. Mentally visualizing and cogitating the whole, the gestalt, the totality of the process is important. I believe this is analogous to hearing a story, to being captured by an story, and dialectic (Plato, not Aristotle, had it right!) Again, during my maritime work and training, I found and find it helpful (often necessary) to be aware of the big-picture of the process, e.g., the goal the reason for the it all, and work through the steps and procedures, in the process.
Respectfully,
Robert Rovetto
rrovetto[at]buffalo[dot]edu (Alumnus)
rrovetto[at]terpalum.umd.edu (Alumnus) (preferred email)
ontologos[at]yahoo[dot]com
Robert Rovetto
rrovetto[at]buffalo[dot]edu (Alumnus)
rrovetto[at]terpalum.umd.edu (Alumnus) (preferred email)
ontologos[at]yahoo[dot]com
--
All contributions to this forum by its members are made under an open content license, open publication license, open source or free software license. Unless otherwise specified, all Ontolog Forum content shall be subject to the Creative Commons CC-BY-SA 4.0 License or its successors.
---
You received this message because you are subscribed to the Google Groups "ontolog-forum" group.
To unsubscribe from this group and stop receiving emails from it, send an email to ontolog-foru...@googlegroups.com.
To post to this group, send email to ontolo...@googlegroups.com.
Visit this group at https://groups.google.com/group/ontolog-forum.
To view this discussion on the web visit https://groups.google.com/d/msgid/ontolog-forum/089af3db-3821-1a99-137b-e888d71a7fa9%40firststarsystems.com.
Reply all
Reply to author
Forward
0 new messages
