New help page for adding metadata to track hubs!
Brian Lee
We're proud to announce a help page for adding metadata to track hubs: http://genome.ucsc.edu/goldenPath/help/metadata.html
The page provides multiple examples of adding metadata for track hubs. Adding metadata to tracks can inform users about cell lines, experimental protocols, or assays for the data being selected and viewed.
The help page is divided into four sections:
- Adding metadata to tracks -providing an introduction
- Previous metadata versions -explaining the historical settings for adding metadata
- tagStorm metadata -explaining a new tagStorm metadata file approach
- Tab-separated metadata -explaining a tab-separated metadata file approach
Metadata has always been possible to add to track hubs in a trackDb.txt file using a metadata setting, as discussed in the "Previous metadata versions" section: http://genome.ucsc.edu/goldenPath/help/metadata.html#metadata
The last two sections of the help page discuss how to add metadata
via the newly supported metaDb (tagStorm) or metaTab (tab-separated)
settings in a track hub's genomes.txt stanza to point to a supplemental
file that contains metadata about each track in a hub. Here's a glimpse
of an example declaration pointing to a metadata file for a hub:
The above uses metaDb to define the location of a tagStorm file,
while below is an example of metaTab pointing to a tab-separated file.
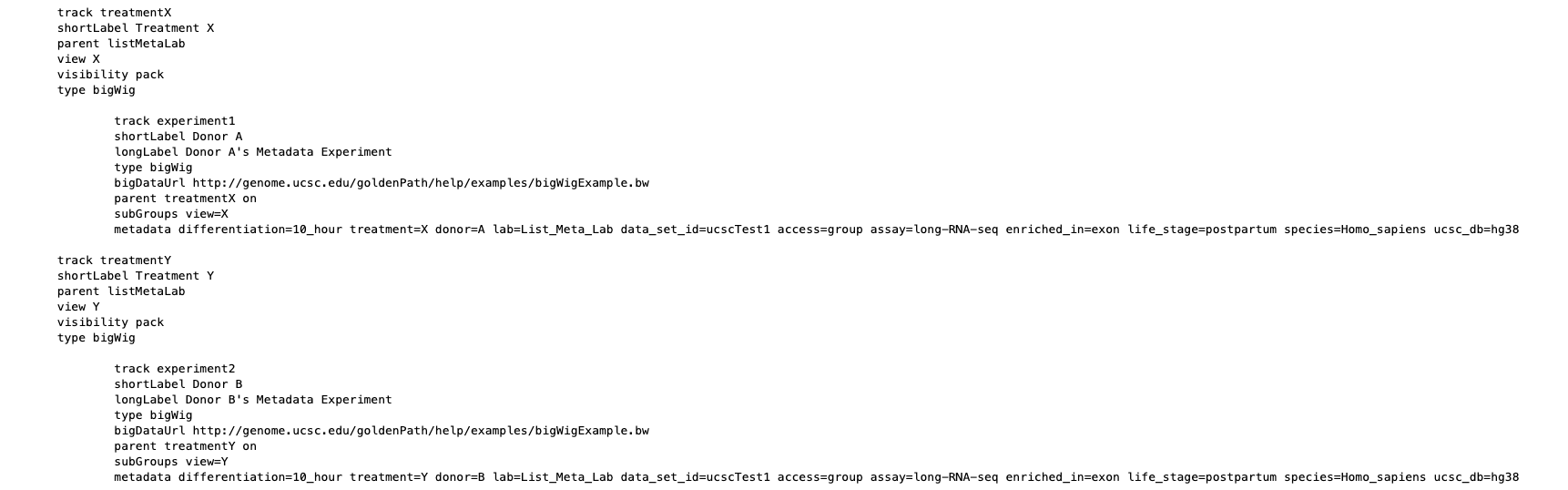
The tagStorm section, https://genome.ucsc.edu/goldenPath/help/metadata.html#tagstorm, demonstrates this new format, which is easy for computers to parse, reduces the redundancy of a tab-separated file, and makes the metadata very human readable. The section includes a canonical tagStorm example showing an experiment of two treatments on three different donors. With tagStorm, each stanza (such as the "donor B" stanza in the example) inherits from any stanzas above it at the right indentation level, and is a parent to stanzas beneath (such as children in the example of different experiment runs with varying differentiation time span). This organization cuts repetition and collapses significant shared information to relevant levels, making the metadata file itself a valuable document to review when understanding a batch of data.
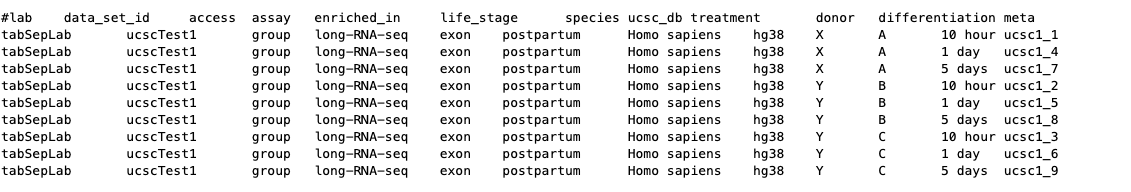
The tab-separated metadata section, http://genome.ucsc.edu/goldenPath/help/metadata.html#tabsep, discusses how column or tab-separated metadata can be used in track hubs and is useful to store information in an array for computer processing. While less human readable than using the tagStorm file approach, adding a TSV or CSV file to a track hub to store metadata is a major improvement over the original implementation, still supported, where each metadata key-value pair must be placed in each stanza of a track that includes metadata, greatly bloating the trackDb.txt file.
Each section of the metadata help page comes with sessions allowing users to quickly click into real interactive demonstration hubs using these different metadata approaches. The example hubs also provide excellent starter templates for building similar hubs.
Example session and hub of original trackDb metaData approach within trackDb.txt: Example session and hub of new additional tagStorm metaData file approach: Example session and hub of new additional tab-separated file metadata approach:Here is the directory for all of these examples: https://genome.ucsc.edu/goldenPath/help/examples/hubExamples/hubMetadata/
Note: All of the above hubs use the "useOneFile on" approach to place the trackDb.txt/genomes.txt/trackDb.txt information in one file: https://genome.ucsc.edu/goldenPath/help/hgTracksHelp.html#UseOneFile
Many thanks to Jairo Navarro and Christopher Lee for building this track hub metadata page, and Braney Raney and Jim Kent for building the tagStorm and tab-separated metadata software.