liftOver Issue
581 views
Skip to first unread message
Christopher Kendall
Apr 23, 2021, 11:57:05 AM4/23/21
to genome...@soe.ucsc.edu
To Whom It May Concern,
I am working with a previously published baboon dataset and I am attempting to use liftOver to convert these from the Panu_2.0 genome to Panu_3.0, and then again to a very new build, Panubis_1.0. I have supplied chain files for 2.0-3.0 and then from 3.0-Panubis1.0
that were given to me by some collaborators, so this dual-stage process is necessary. Unfortunately, I am having an issue with the command line version of liftOver where it keeps erroring out with invalid unsigned integer. This error flag will change
depending on how I have manipulated the files, so it can be any column. I have uploaded the files into R and manually checked that these are not integers, which R confirms. I have rounded the values (screenshot below of what could be considered a float in
column 6) from what could be considered a float to an integer, and this only serves to walk liftOver further along the .bed file and complain again with the invalid unsigned integer error.
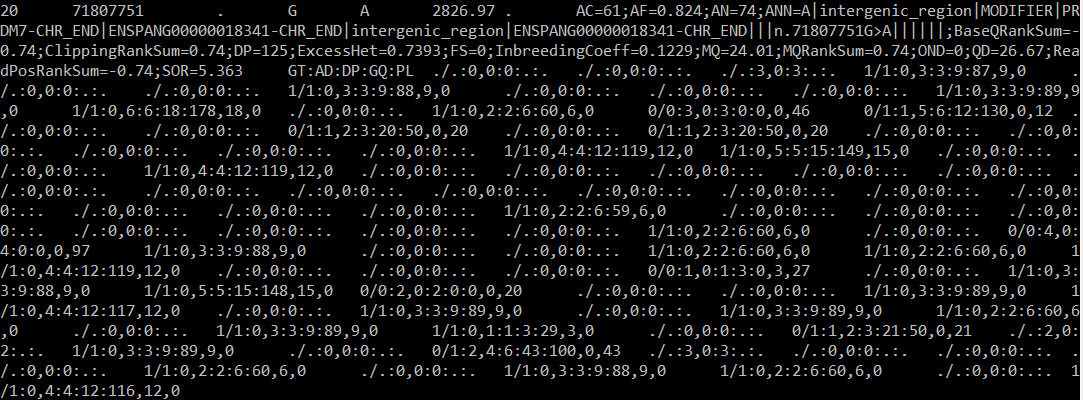
My reasoning for keeping all of the .vcf information is because the Panubis_1.0 build is significantly higher resolution than the previous two builds, which means it would be impossible for me to pinpoint this genomic information in the new build without preserving
these INFO tags if I used a standard .bed file, as some chromosomes change in the new vs. old builds. I have used BEDOPS's VCF2BED command to convert the .vcf files into .bed files along with using the following awk command I found posted somewhere (it might
have even been an archived post here that got a Google hit), and both methods have this error.
grep -v "^#" File.vcf| awk '{printf "%s\t%d\t%d\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\n", $1,$2-1,$2,$3,$4,$5,$6,$7,$8,$9,$10}' > file.bed
The peculiar thing is I have done this in the past using the grep/awk command without issue and used liftOver fine, on this very dataset before without it complaining. Yet now it complains.
My question is: is there a way to preserve all of that VCF information inside of a .bed file with X-number of columns as above and using liftOver without getting thrown this invalid unsigned integer error? I have tried using other programs to circumvent
this such as GATK 4 and their LiftoverVCF and they do work...except for a couple of chromosomes, which cannot be solved at this time. I feel like this invalid unsigned integer error is definitely a new thing as I am following the same workflow as previously
on the same dataset and should be an easy fix I just don't understand how to fix it.
In case you need it, my command and output:
./liftOver Path/to/Bed/Chr20.bed Path/to/Chain/Panu_2.0_Panu_3.0_chr20.chain Path/to/OutputA/test.20.lifted.bed Path/to/OutputB/test.20.unlifted.bed
Reading liftover chains
Mapping coordinates
invalid unsigned integer: "".""
I have had this error throw for the "G", the "A", and 2826.97, all depending on how the columns are shuffled and how I use R to force the columns to be something. It does not seem to be a specific column all the time, which would be helpful.
Thank you for time,
Chris Kendall
Matthew Speir
Apr 26, 2021, 12:45:45 PM4/26/21
to Christopher Kendall, genome...@soe.ucsc.edu
Hello, Chris.
Thank you for your question about LiftOver.
Based on what you have shared about how you are running LiftOver, it sounds like the program is assuming that the columns in your file adhere to the BED standard, https://genome.ucsc.edu/FAQ/FAQformat.html#format1, but it fails because the columns don't contain the values it expects. Since the first handful of columns of the file are standard BED columns, you should use the "-bedPlus=N" option to tell it that only the first N columns of your file are standard and the rest are non-standard. It should preserve those fields past the standard BED fields.
I hope this is helpful. If you have any further questions, please reply to gen...@soe.ucsc.edu. All messages sent to that address are archived on a publicly-accessible Google Groups forum. If your question includes sensitive data, you may send it instead to genom...@soe.ucsc.edu.
Training videos & resources: http://genome.ucsc.edu/training/index.html
Want to share the Browser with colleagues? Host a workshop: http://bit.ly/ucscTraining
Training videos & resources: http://genome.ucsc.edu/training/index.html
Want to share the Browser with colleagues? Host a workshop: http://bit.ly/ucscTraining
---
Matthew Speir
UCSC Cell Browser, Quality Assurance and Data Wrangler
Human Cell Atlas, User Experience Researcher
UCSC Genome Browser, User Support
UC Santa Cruz Genomics Institute
Revealing life’s code.
--
---
You received this message because you are subscribed to the Google Groups "UCSC Genome Browser Mirror-Specific Support" group.
To unsubscribe from this group and stop receiving emails from it, send an email to genome-mirro...@soe.ucsc.edu.
To view this discussion on the web visit https://groups.google.com/a/soe.ucsc.edu/d/msgid/genome-mirror/YT1PR01MB29555582D08252335A6E4719DF459%40YT1PR01MB2955.CANPRD01.PROD.OUTLOOK.COM.
Christopher Kendall
Apr 26, 2021, 1:49:01 PM4/26/21
to Matthew Speir, genome...@soe.ucsc.edu
Hi Matthew,
Thank you! This is insanely helpful and exactly what I was looking for. I appreciate your time.
Sincerely,
Chris
From: Matthew Speir <msp...@ucsc.edu>
Sent: 26 April 2021 12:45
To: Christopher Kendall <chris....@mail.utoronto.ca>
Cc: genome...@soe.ucsc.edu <genome...@soe.ucsc.edu>
Subject: Re: [genome-mirror] liftOver Issue
Sent: 26 April 2021 12:45
To: Christopher Kendall <chris....@mail.utoronto.ca>
Cc: genome...@soe.ucsc.edu <genome...@soe.ucsc.edu>
Subject: Re: [genome-mirror] liftOver Issue
EXTERNAL EMAIL:
Matthew Speir
May 13, 2021, 2:12:50 PM5/13/21
to Christopher Kendall, genome-mirror
Hello, Chris.
Our engineers share that our utilities such as liftOver are, in general, single-thread only (occasionally spawning a child process or two to decompress gzipped input files). The way to achieve parallelization is to reserve only 2 cores (or maybe 4 if multiple inputs are gzipped) and run a lot of separate liftOver jobs at the same time.
I hope this is helpful. If you have any further questions, please reply to genome...@soe.ucsc.edu. All messages sent to that address are archived on a publicly-accessible Google Groups forum. If your question includes sensitive data, you may send it instead to genom...@soe.ucsc.edu.
Training videos & resources: http://genome.ucsc.edu/training/index.html
Want to share the Browser with colleagues? Host a workshop: http://bit.ly/ucscTraining
---
Matthew Speir
UCSC Cell Browser, Quality Assurance and Data Wrangler
Human Cell Atlas, User Experience Researcher
UCSC Genome Browser, User Support
UC Santa Cruz Genomics Institute
Revealing life’s code.
On Thu, May 6, 2021 at 8:18 AM Christopher Kendall <chris....@mail.utoronto.ca> wrote:
Hello,
I had previously sent an email regarding an issue I was having with liftOver, and Matthew Speir was a great help regarding adding -bedPlus=N to my scripts to ignore everything but the first 3 columns in my bed files. I am wondering, as I cannot seem to find the documentation for liftOver command line very readily, is there a command to parallelize the script so it can be hyperthreaded for speed improvement? I am working with a computer cluster that is trying to help me with this but not having much luck. As it stands right now, I am trying to loop over 100 files and run liftOver on each file. However, as it stands, liftOver is using 4MB per task and not using the number of threads I requested, and is instead being a bit of a CPU hog for no reason.
I appreciate the help!
Sincerely,
Chris Kendall
From: Matthew Speir <msp...@ucsc.edu>
Sent: 26 April 2021 12:45
To: Christopher Kendall <chris....@mail.utoronto.ca>
Cc: genome...@soe.ucsc.edu <genome...@soe.ucsc.edu>
Subject: Re: [genome-mirror] liftOver Issue
EXTERNAL EMAIL:
Reply all
Reply to author
Forward
0 new messages
