New gnomAD Mutation Constraint Track
30 views
Skip to first unread message
Daniel Schmelter
May 3, 2022, 7:08:54 PM5/3/22
to genome-...@soe.ucsc.edu
Hello DNA people,
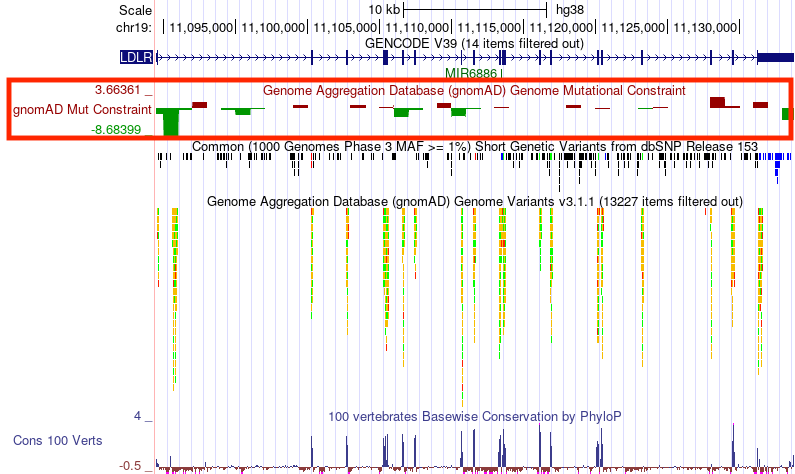
We have just released the GnomAD Genomes Mutation Constraint track on the human GRCh38/hg38 genome assembly. This track's data is based on GnomAD v3.1.2 and shows the relative frequency of variation in 1 kilobase windows across the entire genome. This quantifies the population occurrence of disruptive variation caused by purifying natural selection. This is similar to negative selection on loss-of-function (LoF) for genes but calculated for non-coding regions too. View the complete set of GnomAD tracks and read more on the GnomAD description page.
All the best,
Daniel Schmelter
Reply all
Reply to author
Forward
0 new messages
