question related to add file to my sessions
qiongorwu
Steve Heitner
Hello, Qiong.
When you say that you’re trying to add files to your session, do you mean that you are trying to display additional tracks in your session? Assuming this is the case, after you open the new tracks that you want to view in your session, are you then re-saving the session? If you don’t re-save the session, any time you attempt to load the session again, it will just display whatever tracks were open when you initially saved the session.
Please contact us again at gen...@soe.ucsc.edu if you have any further questions. Questions sent to that address will be archived in a publicly-accessible forum for the benefit of other users. If your question contains sensitive data, you may send it instead to genom...@soe.ucsc.edu.
---
Steve Heitner
UCSC Genome Bioinformatics Group
--
qiongorwu
Matthew Speir
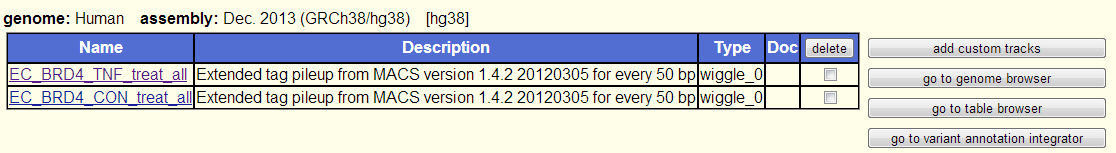
After uploading your new custom tracks, you can re-save your session by going to the "My Data" in the top, blue menu bar and selecting "Sessions". On the following page, if you are logged into your Genome Browser account, you can save new sessions or overwrite your old sessions with new settings.
I hope this is helpful. If you have any further questions, please reply to gen...@soe.ucsc.edu. All messages sent to that address are archived on a publicly-accessible Google Groups forum. If your question includes sensitive data, you may send it instead to genom...@soe.ucsc.edu.
Matthew Speir
UCSC Genome Bioinformatics Group
--
qiongorwu
Jonathan Casper
Hello Qiong,
Your wiggle files for the hg19 genome assembly have coordinates that are specific to hg19. You will need to convert those coordinates into values appropriate for hg18 before you can load your data on that assembly. We do not have a tool to automatically lift wiggle files to new assemblies, but our liftOver tool will work on a related data format: bedGraph. You may be able to convert your wiggle data into bedGraph format, and then use our liftOver tool at http://genome.ucsc.edu/cgi-bin/hgLiftOver to do this conversion. The following mailing list question provides links to some tools that may assist with converting your wiggle data into a bedGraph: https://groups.google.com/a/soe.ucsc.edu/d/topic/genome/fIFzegiIxOw/discussion.
If you prefer, you can also run liftOver on your own computer instead of on our website. This may be necessary if your wiggle data files are particularly large. The liftOver program can be found on our download server at http://hgdownload.soe.ucsc.edu/admin/exe/, already compiled for several computer architectures. If a compiled version for your computer is not available, you may also be able to compile the program yourself from the userApps.src.tgz source code package.
I hope this is helpful. If you have any further questions, please reply to gen...@soe.ucsc.edu or genome...@soe.ucsc.edu. Questions sent to those addresses will be archived in publicly-accessible forums for the benefit of other users. If your question contains sensitive data, you may send it instead to genom...@soe.ucsc.edu.
--
Jonathan Casper
UCSC Genome Bioinformatics Group
--
