BED / narrowPeak : which format to draw this ?
39 views
Skip to first unread message
Benoit Ballester
Oct 2, 2020, 11:27:50 AM10/2/20
to genome...@soe.ucsc.edu
Hi,
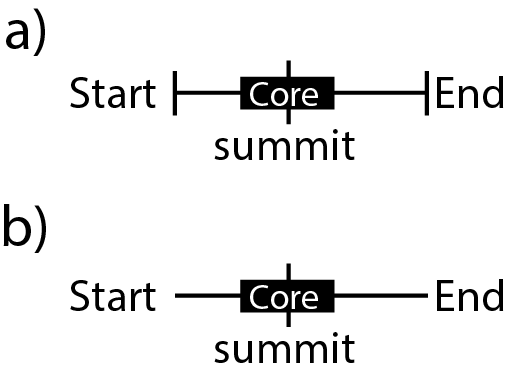
Is there a format that would allow to display data like this in a track (either a or b) ?
I am currently using BED format with thickEnd/thicStart to draw the summit line (for ChIP-seq peaks).
Could anyone advise on a format to get close to this ? Option b) seems to be the most feasible.
Cheers,
Ben
--
Benoît Ballester, PhD
Inserm U1090, TAGC
Campus de Luminy
13288 Marseille Cedex 9
France
Benoît Ballester, PhD
Inserm U1090, TAGC
Campus de Luminy
13288 Marseille Cedex 9
France
Matthew Speir
Oct 6, 2020, 6:09:49 PM10/6/20
to Benoit Ballester, genome-mirror
Hello, Benoit.
Thank you for your question about displaying your data in the UCSC Genome Browser.
Using BED12 format, you should be able to accomplish something similar to what you propose in your Figure A. The BED12 format allows you to define blocks (i.e. exons) and you can combine this with the thickStart/thickEnd to achieve a similar result. In essence, you just need to determine where you want each of the blocks (both outer blocks and the inner block) to start and how large you would like each block to be.
I know that you developed the "ReMap" hubs, including the 2020 version with filtering options. So, here's an example using an item from your track:
track name=original
chrX 15571043 15571512 ENCSR197ALX.HDGF.K-562 0 . 15571367 15571368 112,84,224
track name=bed12_exons
chrX 15571043 15571512 ENCSR197ALX.HDGF.K-562 0 . 15571367 15571368 112,84,224 3 5,200,5 0,205,464
chrX 15571043 15571512 ENCSR197ALX.HDGF.K-562 0 . 15571367 15571368 112,84,224
track name=bed12_exons
chrX 15571043 15571512 ENCSR197ALX.HDGF.K-562 0 . 15571367 15571368 112,84,224 3 5,200,5 0,205,464
You can visualize this in the Genome Browser with this link: http://genome.ucsc.edu/s/brianlee/hg38_customTrack. The 10th column determines the number of blocks, the 11th column the sizes of those blocks, and the 12th column the start point for those blocks relative to the size of the overall element. We did drop the TF and Biotypes columns from our example for ease of copy/paste loading, but you should be able to easily add those back in if you are generating a bigBed track.
I hope this is helpful. If you have any further questions, please reply to genome...@soe.ucsc.edu. All messages sent to that address are archived on a publicly-accessible Google Groups forum. If your question includes sensitive data, you may send it instead to genom...@soe.ucsc.edu.
Training videos & resources: http://genome.ucsc.edu/training/index.html
Want to share the Browser with colleagues? Host a workshop: http://bit.ly/ucscTraining
Training videos & resources: http://genome.ucsc.edu/training/index.html
Want to share the Browser with colleagues? Host a workshop: http://bit.ly/ucscTraining
---
Matthew Speir
UCSC Cell Browser, Quality Assurance and Data Wrangler
Human Cell Atlas, User Experience Researcher
UCSC Genome Browser, User Support
UC Santa Cruz Genomics Institute
Revealing life’s code.
--
---
You received this message because you are subscribed to the Google Groups "UCSC Genome Browser Mirror-Specific Support" group.
To unsubscribe from this group and stop receiving emails from it, send an email to genome-mirro...@soe.ucsc.edu.
To view this discussion on the web visit https://groups.google.com/a/soe.ucsc.edu/d/msgid/genome-mirror/975C2F5A-548C-4BD9-8F40-F168A4B9BA06%40inserm.fr.
Reply all
Reply to author
Forward
0 new messages
